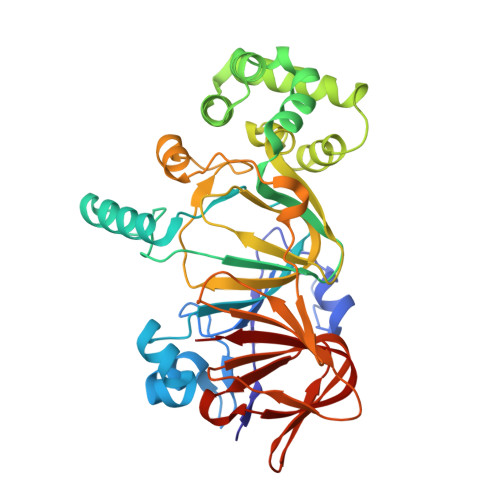
Structural and functional insights into phosphomannose isomerase: the role of zinc and catalytic residues.
Bangera, M., Gowda K, G., Sagurthi, S.R., Murthy, M.R.N.(2019) Acta Crystallogr D Struct Biol 75: 475-487
- PubMed: 31063150 Search on PubMed
- DOI: https://doi.org/10.1107/S2059798319004169
- Primary Citation Related Structures:
5ZT4, 5ZT5, 5ZT6, 5ZUW, 5ZUY, 5ZV0, 5ZVR, 5ZVU, 5ZVX - PubMed Abstract:
Phosphomannose isomerase (PMI) is a housekeeping enzyme that is found in organisms ranging from bacteria to fungi to mammals and is important for cell-wall synthesis, viability and signalling. PMI is a zinc-dependent enzyme that catalyses the reversible isomerization between mannose 6-phosphate (M6P) and fructose 6-phosphate (F6P), presumably via the formation of a cis-enediol intermediate. The reaction is hypothesized to involve ring opening of M6P, the transfer of a proton from the C2 atom to the C1 atom and between the O1 and O2 atoms of the substrate, followed by ring closure resulting in the product F6P. Several attempts have been made to decipher the role of zinc ions and various residues in the catalytic function of PMI. However, there is no consensus on the catalytic base and the mechanism of the reaction catalyzed by the enzyme. In the present study, based on the structure of PMI from Salmonella typhimurium, site-directed mutagenesis targeting residues close to the bound metal ion and activity studies on the mutants, zinc ions were shown to be crucial for substrate binding. These studies also suggest Lys86 as the most probable catalytic base abstracting the proton in the isomerization reaction. Plausible roles for the highly conserved residues Lys132 and Arg274 could also be discerned based on comparison of the crystal structures of wild-type and mutant PMIs. PMIs from prokaryotes possess a low sequence identity to the human enzyme, ranging between 30% and 40%. Since PMI is important for the virulence of many pathogenic organisms, the identification of catalytically important residues will facilitate its use as a potential antimicrobial drug target.
- Molecular Biophysics Unit, Indian Institute of Science, Bangalore, Karnataka 560 012, India.
Organizational Affiliation: