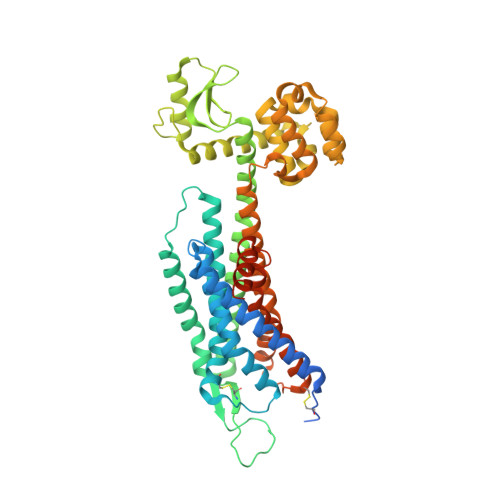
Structural basis of ligand binding modes at the neuropeptide Y Y1receptor
Yang, Z., Han, S., Keller, M., Kaiser, A., Bender, B.J., Bosse, M., Burkert, K., Kogler, L.M., Wifling, D., Bernhardt, G., Plank, N., Littmann, T., Schmidt, P., Yi, C., Li, B., Ye, S., Zhang, R., Xu, B., Larhammar, D., Stevens, R.C., Huster, D., Meiler, J., Zhao, Q., Beck-Sickinger, A.G., Buschauer, A., Wu, B.(2018) Nature 556: 520-524
- PubMed: 29670288 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-018-0046-x
- Primary Citation Related Structures:
5ZBH, 5ZBQ - PubMed Abstract:
Neuropeptide Y (NPY) receptors belong to the G-protein-coupled receptor superfamily and have important roles in food intake, anxiety and cancer biology 1,2 . The NPY-Y receptor system has emerged as one of the most complex networks with three peptide ligands (NPY, peptide YY and pancreatic polypeptide) binding to four receptors in most mammals, namely the Y 1 , Y 2 , Y 4 and Y 5 receptors, with different affinity and selectivity 3 . NPY is the most powerful stimulant of food intake and this effect is primarily mediated by the Y 1 receptor (Y 1 R) 4 . A number of peptides and small-molecule compounds have been characterized as Y 1 R antagonists and have shown clinical potential in the treatment of obesity 4 , tumour 1 and bone loss 5 . However, their clinical usage has been hampered by low potency and selectivity, poor brain penetration ability or lack of oral bioavailability 6 . Here we report crystal structures of the human Y 1 R bound to the two selective antagonists UR-MK299 and BMS-193885 at 2.7 and 3.0 Å resolution, respectively. The structures combined with mutagenesis studies reveal the binding modes of Y 1 R to several structurally diverse antagonists and the determinants of ligand selectivity. The Y 1 R structure and molecular docking of the endogenous agonist NPY, together with nuclear magnetic resonance, photo-crosslinking and functional studies, provide insights into the binding behaviour of the agonist and for the first time, to our knowledge, determine the interaction of its N terminus with the receptor. These insights into Y 1 R can enable structure-based drug discovery that targets NPY receptors.
- Chinese Academy of Sciences Key Laboratory of Receptor Research, Shanghai Institute of Materia Medica, Chinese Academy of Sciences, Shanghai, China.
Organizational Affiliation: