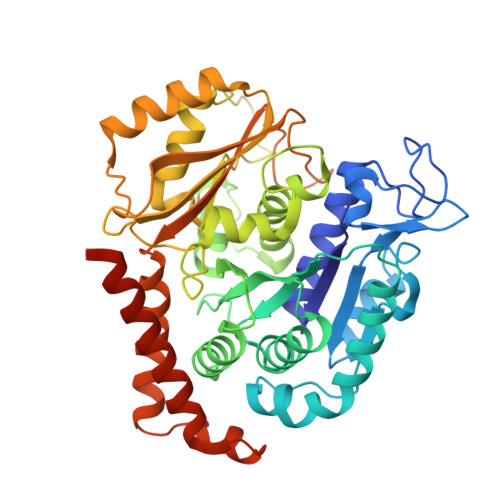
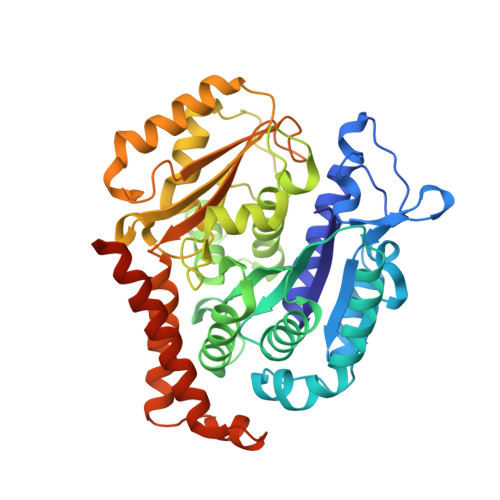
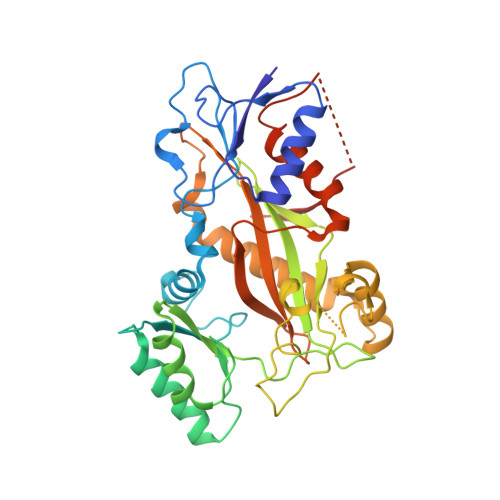
The compound millepachine and its derivatives inhibit tubulin polymerization by irreversibly binding to the colchicine-binding site in beta-tubulin.
Yang, J.H., Yan, W., Yu, Y.M., Wang, Y.X., Yang, T., Xue, L.L., Yuan, X., Long, C.F., Liu, Z.W., Chen, X.X., Hu, M.S., Zheng, L., Qiu, Q., Pei, H.Y., Li, D., Wang, F., Bai, P., Wen, J.L., Ye, H.Y., Chen, L.J.(2018) J Biological Chem 293: 9461-9472
- PubMed: 29691282 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA117.001658
- Primary Citation Related Structures:
5XIW, 5XP3, 5YL2, 5YLJ, 5YLS - PubMed Abstract:
Inhibitors that bind to the paclitaxel- or vinblastine-binding sites of tubulin have been part of the pharmacopoeia of anticancer therapy for decades. However, tubulin inhibitors that bind to the colchicine-binding site are not used in clinical cancer therapy, because of their low therapeutic index. To address multidrug resistance to many conventional tubulin-binding agents, numerous efforts have attempted to clinically develop inhibitors that bind the colchicine-binding site. Previously, we have found that millepachine (MIL), a natural chalcone-type small molecule extracted from the plant Millettia pachycarpa , and its two derivatives (MDs) SKLB028 and SKLB050 have potential antitumor activities both in vitro and in vivo However, their cellular targets and mechanisms are unclear. Here, biochemical and cellular experiments revealed that the MDs directly and irreversibly bind β-tubulin. X-ray crystallography of the tubulin-MD structures disclosed that the MDs bind at the tubulin intradimer interface and to the same site as colchicine and that their binding mode is similar to that of colchicine. Of note, MDs inhibited tubulin polymerization and caused G 2 /M cell-cycle arrest. Comprehensive analysis further revealed that free MIL exhibits an s- cis conformation, whereas MIL in the colchicine-binding site in tubulin adopts an s- trans conformation. Moreover, introducing an α-methyl to MDs to increase the proportion of s- trans conformations augmented MDs' tubulin inhibition activity. Our study uncovers a new class of chalcone-type tubulin inhibitors that bind the colchicine-binding site in β-tubulin and suggests that the s- trans conformation of these compounds may make them more active anticancer agents.
- From the State Key Laboratory of Biotherapy and Cancer Center, West China Hospital, Sichuan University, and Collaborative Innovation Center for Biotherapy, Chengdu 610041, China and.
Organizational Affiliation: