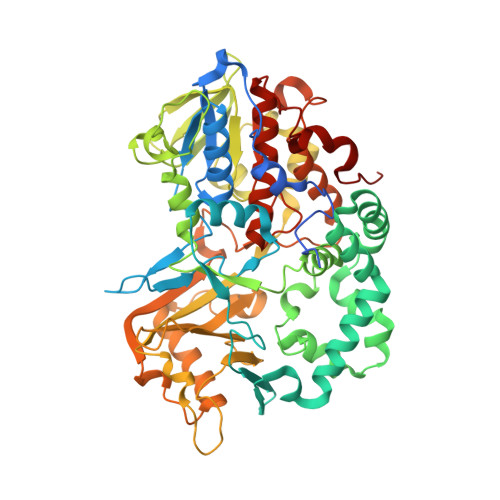
Ligand complex structures of l-amino acid oxidase/monooxygenase from
Im, D., Matsui, D., Arakawa, T., Isobe, K., Asano, Y., Fushinobu, S.(2018) FEBS Open Bio 8: 314-324
- PubMed: 29511608 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/2211-5463.12387
- Primary Citation Related Structures:
5YB6, 5YB7, 5YB8 - PubMed Abstract:
l-Amino acid oxidase/monooxygenase from Pseudomonas sp. AIU 813 (l-AAO/MOG) catalyzes both the oxidative deamination and oxidative decarboxylation of the α-group of l-Lys to produce a keto acid and amide, respectively. l-AAO/MOG exhibits limited specificity for l-amino acid substrates with a basic side chain. We previously determined its ligand-free crystal structure and identified a key residue for maintaining the dual activities. Here, we determined the structures of l-AAO/MOG complexed with l-Lys, l-ornithine, and l-Arg and revealed its substrate recognition. Asp238 is located at the ceiling of a long hydrophobic pocket and forms a strong interaction with the terminal, positively charged group of the substrates. A mutational analysis on the D238A mutant indicated that the interaction is critical for substrate binding but not for catalytic control between the oxidase/monooxygenase activities. The catalytic activities of the D238E mutant unexpectedly increased, while the D238F mutant exhibited altered substrate specificity to long hydrophobic substrates. In the ligand-free structure, there are two channels connecting the active site and solvent, and a short region located at the dimer interface is disordered. In the l-Lys complex structure, a loop region is displaced to plug the channels. Moreover, the disordered region in the ligand-free structure forms a short helix in the substrate complex structures and creates the second binding site for the substrate. It is assumed that the amino acid substrate enters the active site of l-AAO/MOG through this route. The atomic coordinates and structure factors (codes 5YB6, 5YB7, and 5YB8) have been deposited in the Protein Data Bank (http://wwpdb.org/). 1.4.3.2 (l-amino acid oxidase), 1.13.12.2 (lysine 2-monooxygenase).
- Department of Biotechnology The University of Tokyo Japan.
Organizational Affiliation: