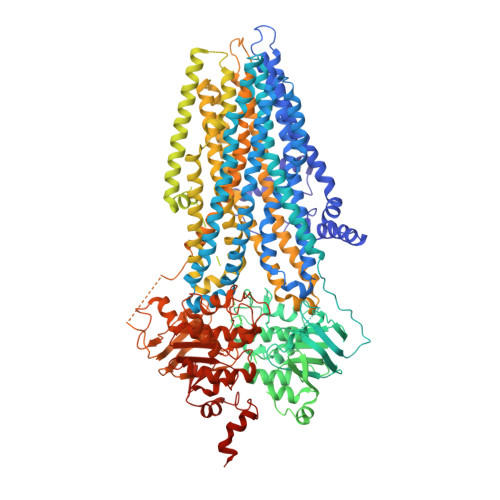
Conformational Changes of CFTR upon Phosphorylation and ATP Binding.
Zhang, Z., Liu, F., Chen, J.(2017) Cell 170: 483-491.e8
- PubMed: 28735752 Search on PubMed
- DOI: https://doi.org/10.1016/j.cell.2017.06.041
- Primary Citation Related Structures:
5W81 - PubMed Abstract:
The cystic fibrosis transmembrane conductance regulator (CFTR) is an anion channel evolved from an ATP-binding cassette transporter. CFTR channel gating is strictly coupled to phosphorylation and ATP hydrolysis. Previously, we reported essentially identical structures of zebrafish and human CFTR in the dephosphorylated, ATP-free form. Here, we present the structure of zebrafish CFTR in the phosphorylated, ATP-bound conformation, determined by cryoelectron microscopy to 3.4 Å resolution. Comparison of the two conformations shows major structural rearrangements leading to channel opening. The phosphorylated regulatory domain is disengaged from its inhibitory position; the nucleotide-binding domains (NBDs) form a "head-to-tail" dimer upon binding ATP; and the cytoplasmic pathway, found closed off in other ATP-binding cassette transporters, is cracked open, consistent with CFTR's unique channel function. Unexpectedly, the extracellular mouth of the ion pore remains closed, indicating that local movements of the transmembrane helices can control ion access to the pore even in the NBD-dimerized conformation.
- Laboratory of Membrane Biophysics and Biology, The Rockefeller University, New York, NY, USA; Howard Hughes Medical Institute, Chevy Chase, MD 20815, USA.
Organizational Affiliation: