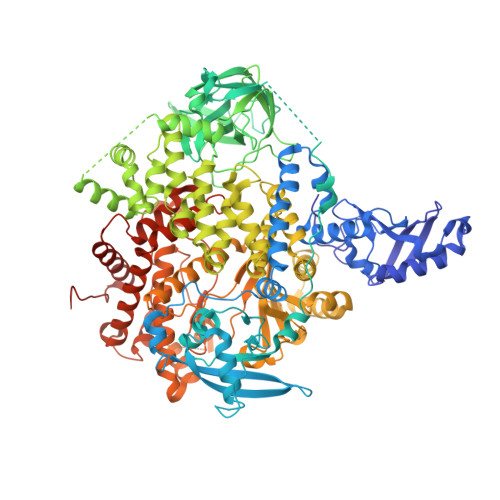
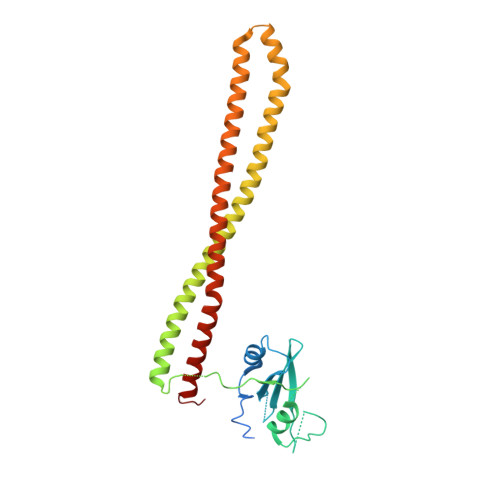
Rationally Designed PI3K alpha Mutants to Mimic ATR and Their Use to Understand Binding Specificity of ATR Inhibitors.
Lu, Y., Knapp, M., Crawford, K., Warne, R., Elling, R., Yan, K., Doyle, M., Pardee, G., Zhang, L., Ma, S., Mamo, M., Ornelas, E., Pan, Y., Bussiere, D., Jansen, J., Zaror, I., Lai, A., Barsanti, P., Sim, J.(2017) J Mol Biology 429: 1684-1704
- PubMed: 28433539 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2017.04.006
- Primary Citation Related Structures:
5UK8, 5UKJ, 5UL1 - PubMed Abstract:
ATR, a protein kinase in the PIKK family, plays a critical role in the cell DNA-damage response and is an attractive anticancer drug target. Several potent and selective inhibitors of ATR have been reported showing significant antitumor efficacy, with most advanced ones entering clinical trials. However, due to the absence of an experimental ATR structure, the determinants contributing to ATR inhibitors' potency and specificity are not well understood. Here we present the mutations in the ATP-binding site of PI3Kα to progressively transform the pocket to mimic that of ATR. The generated PI3Kα mutants exhibit significantly improved affinity for selective ATR inhibitors in multiple chemical classes. Furthermore, we obtained the X-ray structures of the PI3Kα mutants in complex with the ATR inhibitors. The crystal structures together with the analysis on the inhibitor affinity profile elucidate the roles of individual amino acid residues in the binding of ATR inhibitors, offering key insights for the binding mechanism and revealing the structure features important for the specificity of ATR inhibitors. The ability to obtain structural and binding data for these PI3Kα mutants, together with their ATR-like inhibitor binding profiles, makes these chimeric PI3Kα proteins valuable model systems for structure-based inhibitor design.
- Global Discovery Chemistry, Novartis Institutes for Biomedical Research, Emeryville, CA 94608, USA. Electronic address: yipin.lu@novartis.com.
Organizational Affiliation: