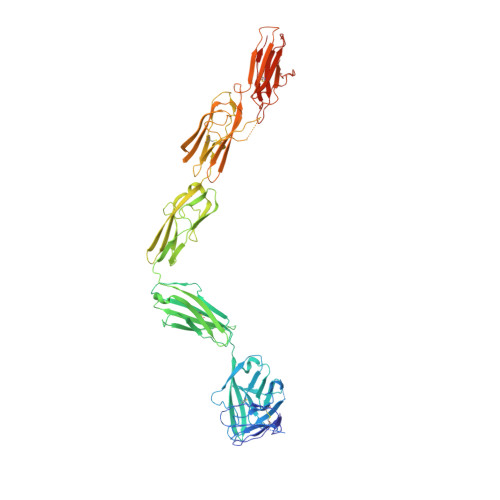
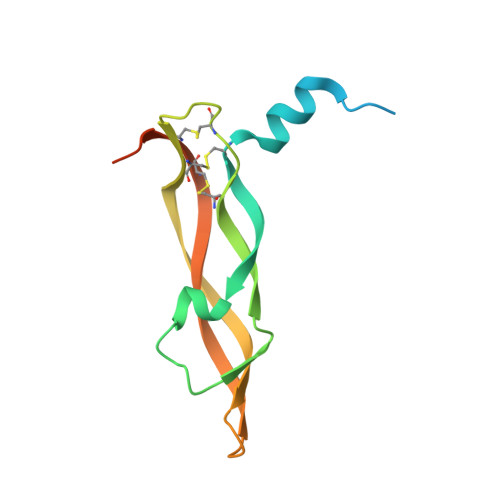
Structure of the Full-length VEGFR-1 Extracellular Domain in Complex with VEGF-A.
Markovic-Mueller, S., Stuttfeld, E., Asthana, M., Weinert, T., Bliven, S., Goldie, K.N., Kisko, K., Capitani, G., Ballmer-Hofer, K.(2017) Structure 25: 341-352
- PubMed: 28111021 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2016.12.012
- Primary Citation Related Structures:
5T89 - PubMed Abstract:
Vascular endothelial growth factors (VEGFs) regulate blood and lymph vessel development upon activation of three receptor tyrosine kinases: VEGFR-1, -2, and -3. Partial structures of VEGFR/VEGF complexes based on single-particle electron microscopy, small-angle X-ray scattering, and X-ray crystallography revealed the location of VEGF binding and domain arrangement of individual receptor subdomains. Here, we describe the structure of the full-length VEGFR-1 extracellular domain in complex with VEGF-A at 4 Å resolution. We combined X-ray crystallography, single-particle electron microscopy, and molecular modeling for structure determination and validation. The structure reveals the molecular details of ligand-induced receptor dimerization, in particular of homotypic receptor interactions in immunoglobulin homology domains 4, 5, and 7. Functional analyses of ligand binding and receptor activation confirm the relevance of these homotypic contacts and identify them as potential therapeutic sites to allosterically inhibit VEGFR-1 activity.
- Paul Scherrer Institute, Laboratory of Biomolecular Research, 5232 Villigen, Switzerland.
Organizational Affiliation: