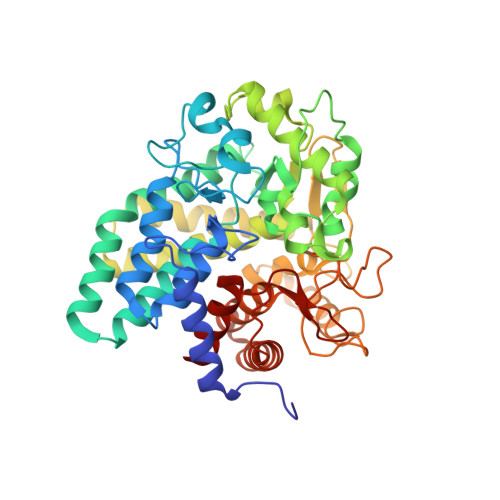
The structural characterization of a glucosylglycerate hydrolase provides insights into the molecular mechanism of mycobacterial recovery from nitrogen starvation.
Cereija, T.B., Alarico, S., Lourenco, E.C., Manso, J.A., Ventura, M.R., Empadinhas, N., Macedo-Ribeiro, S., Pereira, P.J.B.(2019) IUCrJ 6: 572-585
- PubMed: 31316802 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2052252519005372
- Primary Citation Related Structures:
5OHC, 5OHZ, 5OI0, 5OI1, 5OIE, 5OIV, 5OIW, 5OJ4, 5OJU, 5OJV, 5ONT, 5ONZ, 5OO2, 6Q5T - PubMed Abstract:
Bacteria are challenged to adapt to environmental variations in order to survive. Under nutritional stress, several bacteria are able to slow down their metabolism into a nonreplicating state and wait for favourable conditions. It is almost universal that bacteria accumulate carbon stores to survive during this nonreplicating state and to fuel rapid proliferation when the growth-limiting stress disappears. Mycobacteria are exceedingly successful in their ability to become dormant under harsh circumstances and to be able to resume growth when conditions are favourable. Rapidly growing mycobacteria accumulate glucosylglycerate under nitrogen-limiting conditions and quickly mobilize it when nitrogen availability is restored. The depletion of intracellular glucosyl-glycerate levels in Mycolicibacterium hassiacum (basonym Mycobacterium hassiacum ) was associated with the up-regulation of the gene coding for glucosylglycerate hydrolase (GgH), an enzyme that is able to hydrolyse glucosylglycerate to glycerate and glucose, a source of readily available energy. Highly conserved among unrelated phyla, GgH is likely to be involved in bacterial reactivation following nitrogen starvation, which in addition to other factors driving mycobacterial recovery may also provide an opportunity for therapeutic intervention, especially in the serious infections caused by some emerging opportunistic pathogens of this group, such as Mycobacteroides abscessus (basonym Mycobacterium abscessus ). Using a combination of biochemical methods and hybrid structural approaches, the oligomeric organization of M. hassiacum GgH was determined and molecular determinants of its substrate binding and specificity were unveiled.
- IBMC - Instituto de Biologia Molecular e Celular, Universidade do Porto, Porto, Portugal.
Organizational Affiliation: