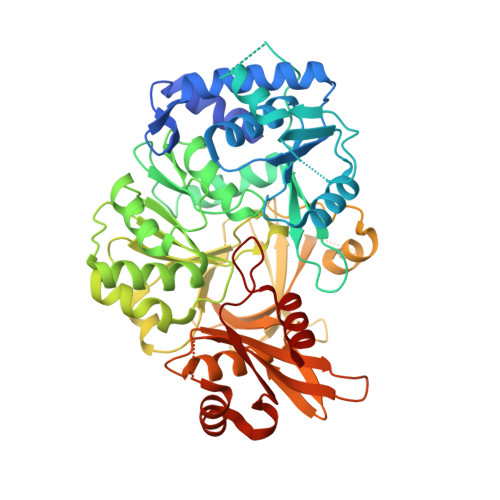
Structure of the adenylation domain Thr1 involved in the biosynthesis of 4-chlorothreonine in Streptomyces sp. OH-5093-protein flexibility and molecular bases of substrate specificity.
Scaglione, A., Fullone, M.R., Montemiglio, L.C., Parisi, G., Zamparelli, C., Vallone, B., Savino, C., Grgurina, I.(2017) FEBS J 284: 2981-2999
- PubMed: 28704585 Search on PubMed
- DOI: https://doi.org/10.1111/febs.14163
- Primary Citation Related Structures:
5N9W, 5N9X - PubMed Abstract:
We determined the crystal structure of Thr1, the self-standing adenylation domain involved in the nonribosomal-like biosynthesis of free 4-chlorothreonine in Streptomyces sp. OH-5093. Thr1 shows two monomers in the crystallographic asymmetric unit with different relative orientations of the C- and N-terminal subdomains both in the presence of substrates and in the unliganded form. Cocrystallization with substrates, adenosine 5'-triphosphate and l-threonine, yielded one monomer containing the two substrates and the other in complex with l-threonine adenylate, locked in a postadenylation state. Steady-state kinetics showed that Thr1 activates l-Thr and its stereoisomers, as well as d-Ala, l- and d-Ser, albeit with lower efficiency. Modeling of these substrates in the active site highlighted the molecular bases of substrate discrimination. This work provides the first crystal structure of a threonine-activating adenylation enzyme, a contribution to the studies on conformational rearrangement in adenylation domains and on substrate recognition in nonribosomal biosynthesis. Structural data are available in the Protein Data Bank under the accession number 5N9W and 5N9X.
- Department of Biochemical Sciences "A. Rossi Fanelli", Istituto Pasteur-Fondazione Cenci Bolognetti, Sapienza University of Rome, Italy.
Organizational Affiliation: