(131)I-labeled Anti-HER2 Camelid sdAb as a Theranostic Tool in Cancer Treatment.
D'Huyvetter, M., De Vos, J., Xavier, C., Pruszynski, M., Sterckx, Y.G.J., Massa, S., Raes, G., Caveliers, V., Zalutsky, M.R., Lahoutte, T., Devoogdt, N.(2017) Clin Cancer Res 23: 6616-6628
- PubMed: 28751451 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1158/1078-0432.CCR-17-0310
- Primary Citation Related Structures:
5MY6 - PubMed Abstract:
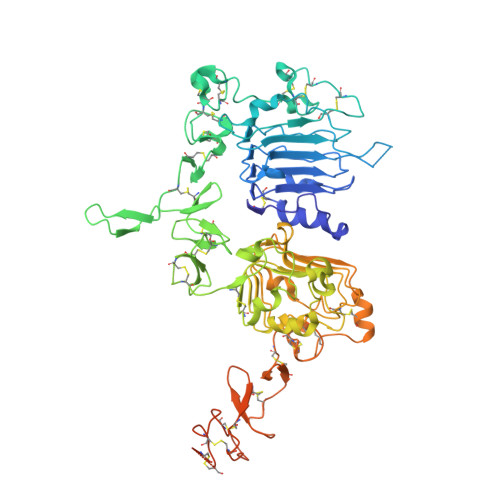
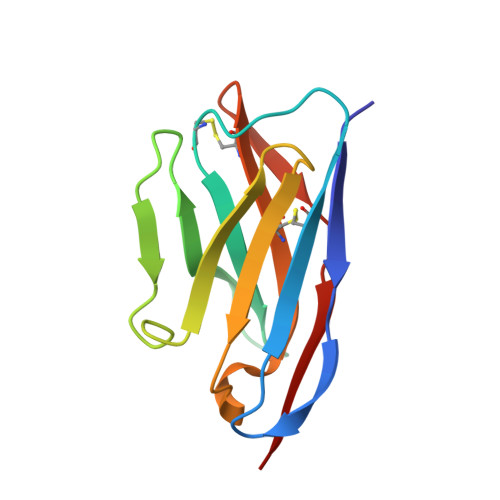
Purpose: Camelid single-domain antibody-fragments (sdAb) have beneficial pharmacokinetic properties, and those targeted to HER2 can be used for imaging of HER2-overexpressing cancer. Labeled with a therapeutic radionuclide, they may be used for HER2-targeted therapy. Here, we describe the generation of a 131 I-labeled sdAb as a theranostic drug to treat HER2-overexpressing cancer. Experimental Design: Anti-HER2 sdAb 2Rs15d was labeled with 131 I using [ 131 I]SGMIB and evaluated in vitro Biodistribution was evaluated in two HER2 + murine xenograft models by micro-SPECT/CT imaging and at necropsy, and under challenge with trastuzumab and pertuzumab. The therapeutic potential of [ 131 I]SGMIB-2Rs15d was investigated in two HER2 + tumor mouse models. A single-dose toxicity study was performed in mice using unlabeled [ 127 I]SGMIB-sdAb at 1.4 mg/kg. The structure of the 2Rs15d-HER2 complex was determined by X-ray crystallography. Results: [ 131 I]SGMIB-2Rs15d bound specifically to HER2 + cells ( K d = 4.74 ± 0.39 nmol/L). High and specific tumor uptake was observed in both BT474/M1 and SKOV-3 tumor xenografted mice and surpassed kidney levels by 3 hours. Extremely low uptake values were observed in other normal tissues at all time points. The crystal structure revealed that 2Rs15d recognizes HER2 Domain 1, consistent with the lack of competition with trastuzumab and pertuzumab observed in vivo [ 131 I]SGMIB-2Rs15d alone, or in combination with trastuzumab, extended median survival significantly. No toxicity was observed after injecting [ 127 I]SGMIB-2Rs15d. Conclusions: These findings demonstrate the theranostic potential of [ 131 I]SGMIB-2Rs15d. An initial scan using low radioactive [*I]SGMIB-2Rs15d allows patient selection and dosimetry calculations for subsequent therapeutic [ 131 I]SGMIB-2Rs15d and could thereby impact therapy outcome on HER2 + breast cancer patients. Clin Cancer Res; 23(21); 6616-28. ©2017 AACR .
- In Vivo Cellular and Molecular Imaging Laboratory, Vrije Universiteit Brussel, Brussels, Belgium. mdhuyvet@vub.ac.be.
Organizational Affiliation: