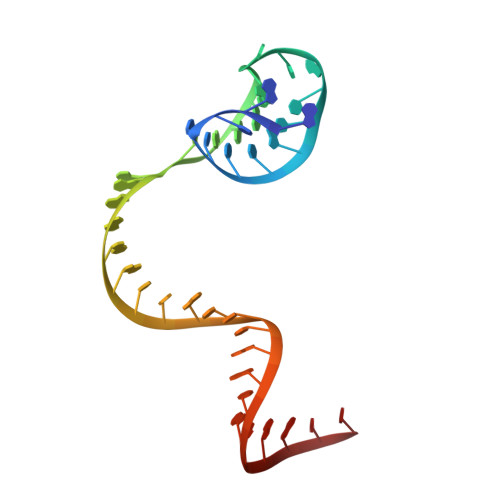
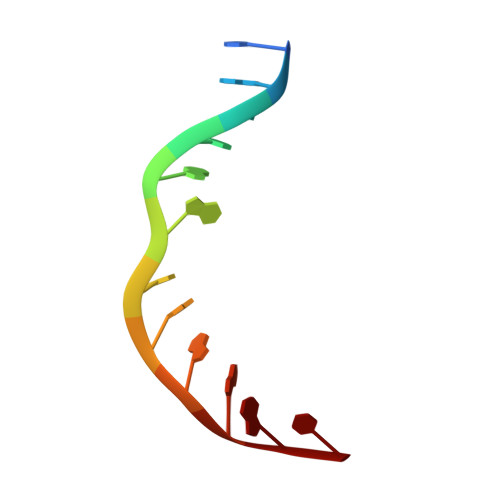
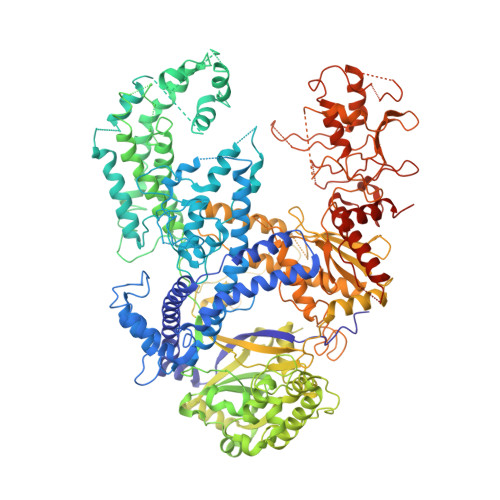
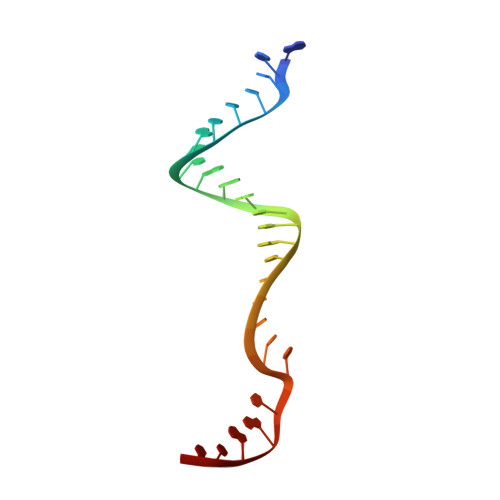
Structure of the Cpf1 endonuclease R-loop complex after target DNA cleavage.
Stella, S., Alcon, P., Montoya, G.(2017) Nature 546: 559-563
- PubMed: 28562584 Search on PubMed
- DOI: https://doi.org/10.1038/nature22398
- Primary Citation Related Structures:
5MGA - PubMed Abstract:
Cpf1 is an RNA-guided endonuclease that is emerging as a powerful genome-editing tool. Here we provide insight into its DNA-targeting mechanism by determining the structure of Francisella novicida Cpf1 with the triple-stranded R-loop generated after DNA cleavage. The structure reveals the machinery involved in DNA unwinding to form a CRISPR RNA (crRNA)-DNA hybrid and a displaced DNA strand. The protospacer adjacent motif (PAM) is recognized by the PAM-interacting domain. The loop-lysine helix-loop motif in this domain contains three conserved lysine residues that are inserted in a dentate manner into the double-stranded DNA. Unzipping of the double-stranded DNA occurs in a cleft arranged by acidic and hydrophobic residues facilitating the crRNA-DNA hybrid formation. The PAM single-stranded DNA is funnelled towards the nuclease site through a mixed hydrophobic and basic cavity. In this catalytic conformation, the PAM-interacting domain and the helix-loop-helix motif in the REC1 domain adopt a 'rail' shape and 'flap-on' conformations, respectively, channelling the PAM strand into the cavity. A steric barrier between the RuvC-II and REC1 domains forms the 'septum', separating the displaced PAM strand and the crRNA-DNA hybrid, avoiding DNA re-annealing. Mutations in key residues reveal a mechanism linking the PAM and DNA nuclease sites. Analysis of the Cpf1 structures proposes a singular working model of RNA-guided DNA cleavage, suggesting new avenues for redesign of Cpf1.
- Protein Structure &Function Programme, Macromolecular Crystallography Group, Novo Nordisk Foundation Center for Protein Research, Faculty of Health and Medical Sciences, University of Copenhagen, Blegdamsvej 3B, Copenhagen 2200, Denmark.
Organizational Affiliation: