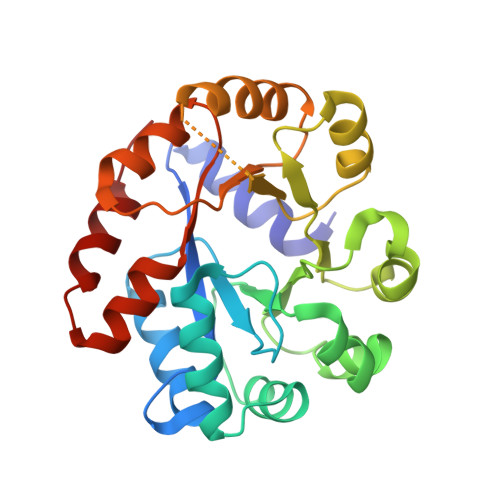
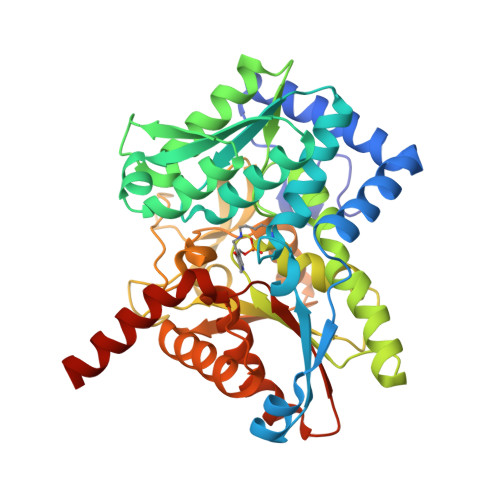
Conservation of the structure and function of bacterial tryptophan synthases.
Michalska, K., Gale, J., Joachimiak, G., Chang, C., Hatzos-Skintges, C., Nocek, B., Johnston, S.E., Bigelow, L., Bajrami, B., Jedrzejczak, R.P., Wellington, S., Hung, D.T., Nag, P.P., Fisher, S.L., Endres, M., Joachimiak, A.(2019) IUCrJ 6: 649-664
- PubMed: 31316809 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2052252519005955
- Primary Citation Related Structures:
5KIN, 5KZM - PubMed Abstract:
Tryptophan biosynthesis is one of the most characterized processes in bacteria, in which the enzymes from Salmonella typhimurium and Escherichia coli serve as model systems. Tryptophan synthase (TrpAB) catalyzes the final two steps of tryptophan biosynthesis in plants, fungi and bacteria. This pyridoxal 5'-phosphate (PLP)-dependent enzyme consists of two protein chains, α (TrpA) and β (TrpB), functioning as a linear αββα heterotetrameric complex containing two TrpAB units. The reaction has a complicated, multistep mechanism resulting in the β-replacement of the hydroxyl group of l-serine with an indole moiety. Recent studies have shown that functional TrpAB is required for the survival of pathogenic bacteria in macrophages and for evading host defense. Therefore, TrpAB is a promising target for drug discovery, as its orthologs include enzymes from the important human pathogens Streptococcus pneumoniae , Legionella pneumophila and Francisella tularensis , the causative agents of pneumonia, legionnaires' disease and tularemia, respectively. However, specific biochemical and structural properties of the TrpABs from these organisms have not been investigated. To fill the important phylogenetic gaps in the understanding of TrpABs and to uncover unique features of TrpAB orthologs to spearhead future drug-discovery efforts, the TrpABs from L. pneumophila , F. tularensis and S. pneumoniae have been characterized. In addition to kinetic properties and inhibitor-sensitivity data, structural information gathered using X-ray crystallo-graphy is presented. The enzymes show remarkable structural conservation, but at the same time display local differences in both their catalytic and allosteric sites that may be responsible for the observed differences in catalysis and inhibitor binding. This functional dissimilarity may be exploited in the design of species-specific enzyme inhibitors.
- Center for Structural Genomics of Infectious Diseases, University of Chicago, Chicago, IL 60367, USA.
Organizational Affiliation: