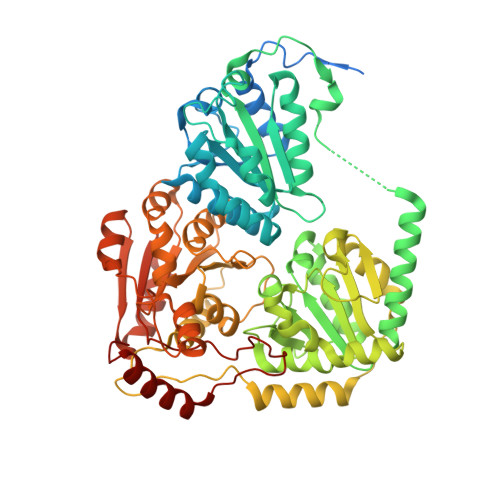
Commercial Herbicides Can Trigger the Oxidative Inactivation of Acetohydroxyacid Synthase.
Lonhienne, T., Nouwens, A., Williams, C.M., Fraser, J.A., Lee, Y.T., West, N.P., Guddat, L.W.(2016) Angew Chem Int Ed Engl 55: 4247-4251
- PubMed: 26924714 Search on PubMed
- DOI: https://doi.org/10.1002/anie.201511985
- Primary Citation Related Structures:
5FEM - PubMed Abstract:
Acetohydroxyacid synthase (AHAS) inhibitors are highly successful commercial herbicides. New kinetic data show that the binding of these compounds leads to reversible accumulative inhibition of AHAS. Crystallographic data (to a resolution of 2.17 Å) for an AHAS-herbicide complex shows that closure of the active site occurs when the herbicidal inhibitor binds, thus preventing exchange with solvent. This feature combined with new kinetic data shows that molecular oxygen promotes an accumulative inhibition leading to the conclusion that the exceptional potency of these herbicides is augmented by subversion of an inherent oxygenase side reaction. The reactive oxygen species produced by this reaction are trapped in the active site, triggering oxidation reactions that ultimately lead to the alteration of the redox state of the cofactor flavin adenine dinucleotide (FAD), a feature that accounts for the observed reversible accumulative inhibition.
- School of Chemistry and Molecular Biosciences, The University of Queensland, Brisbane, 4072 QLD, Australia. t.lonhienne@uq.edu.au.
Organizational Affiliation: