Comprehensive analysis of antibody recognition in convalescent humans from highly pathogenic avian influenza H5N1 infection
Zuo, T., Sun, J., Wang, G., Jiang, L., Zuo, Y., Li, D., Shi, X., Liu, X., Fan, S., Ren, H., Hu, H., Sun, L., Zhou, B., Liang, M., Zhou, P., Wang, X., Zhang, L.(2015) Nat Commun 6: 8855-8855
- PubMed: 26635249 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms9855
- Primary Citation Related Structures:
5DUM, 5DUP, 5DUR, 5DUT - PubMed Abstract:
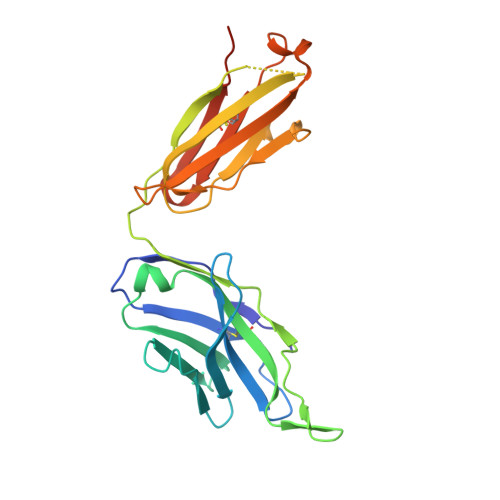
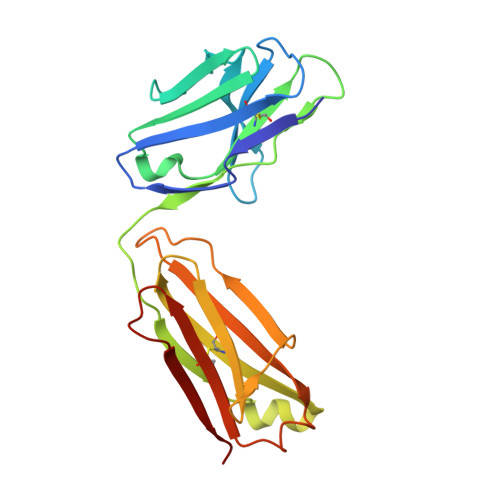
Understanding the mechanism of protective antibody recognition against highly pathogenic avian influenza A virus H5N1 in humans is critical for the development of effective therapies and vaccines. Here we report the crystal structure of three H5-specific human monoclonal antibodies bound to the globular head of hemagglutinin (HA) with distinct epitope specificities, neutralization potencies and breadth. A structural and functional analysis of these epitopes combined with those reported elsewhere identifies four major vulnerable sites on the globular head of H5N1 HA. Chimeric and vulnerable site-specific mutant pseudoviruses are generated to delineate broad neutralization specificities of convalescent sera from two individuals who recovered from the infection with H5N1 virus. Our results show that the four vulnerable sites on the globular head rather than the stem region are the major neutralizing targets, suggesting that during natural H5N1 infection neutralizing antibodies against the globular head work in concert to provide protective antibody-mediated immunity.
- Comprehensive AIDS Research Center, Collaborative Innovation Center for Diagnosis and Treatment of Infectious Diseases, School of Medicine, Tsinghua University, Beijing 100084, China.
Organizational Affiliation: