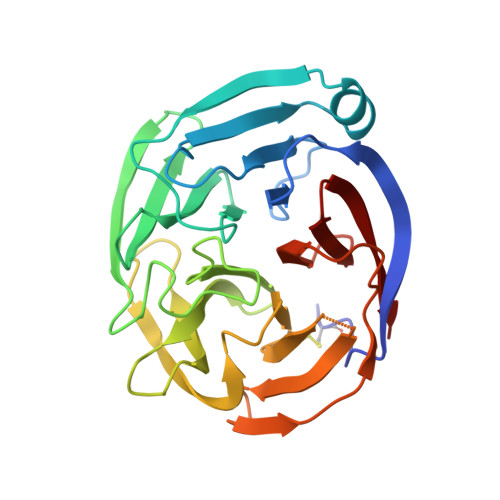
Structural Basis of Latrophilin-FLRT-UNC5 Interaction in Cell Adhesion.
Lu, Y.C., Nazarko, O.V., Sando, R., Salzman, G.S., Sudhof, T.C., Arac, D.(2015) Structure 23: 1678-1691
- PubMed: 26235030 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2015.06.024
- Primary Citation Related Structures:
5CMN, 5CMP - PubMed Abstract:
Fibronectin leucine-rich repeat transmembrane proteins (FLRTs) are cell-adhesion molecules with emerging functions in cortical development and synapse formation. Their extracellular regions interact with latrophilins (LPHNs) to mediate synapse development, and with Uncoordinated-5 (UNC5)/netrin receptors to control the migration of neurons in the developing cortex. Here, we present the crystal structures of FLRT3 in isolation and in complex with LPHN3. The LPHN3/FLRT3 structure reveals that LPHN3 binds to FLRT3 at a site distinct from UNC5. Structure-based mutations specifically disrupt LPHN3/FLRT3 binding, but do not disturb their interactions with other proteins or their cell-membrane localization. Thus, they can be used as molecular tools to dissect the functions of FLRTs and LPHNs in vivo. Our results suggest that UNC5 and LPHN3 can simultaneously bind to FLRT3, forming a trimeric complex, and that FLRT3 may form transsynaptic complexes with both LPHN3 and UNC5. These findings provide molecular insights for understanding the role of cell-adhesion proteins in synapse function.
- Department of Biochemistry and Molecular Biology, The University of Chicago, 929 East 57(th) Street, GCIS W219, Chicago, IL 60637, USA.
Organizational Affiliation: