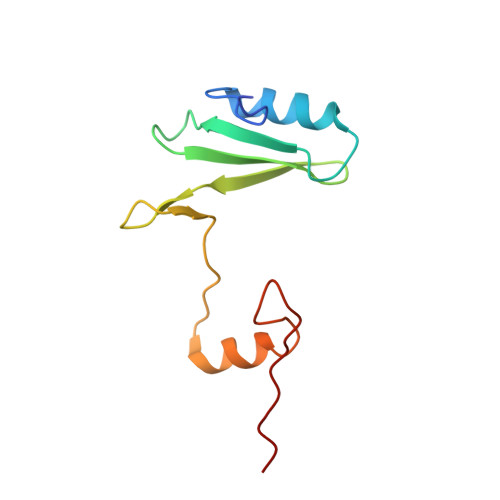
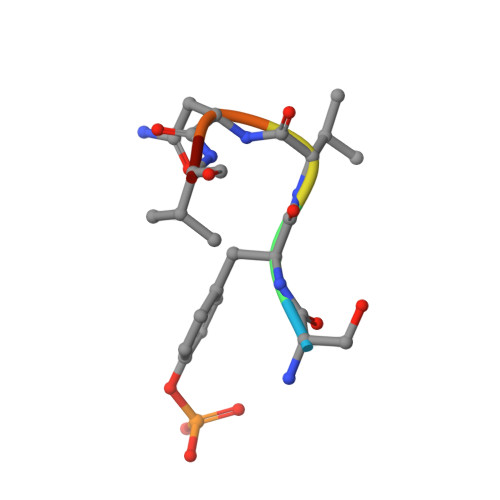
Structural and biophysical investigation of the interaction of a mutant Grb2 SH2 domain (W121G) with its cognate phosphopeptide.
Papaioannou, D., Geibel, S., Kunze, M.B., Kay, C.W., Waksman, G.(2016) Protein Sci 25: 627-637
- PubMed: 26645482 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.2856
- Primary Citation Related Structures:
5CDW - PubMed Abstract:
The adaptor protein Grb2 is a key element of mitogenetically important signaling pathways. With its SH2 domain it binds to upstream targets while its SH3 domains bind to downstream proteins thereby relaying signals from the cell membranes to the nucleus. The Grb2 SH2 domain binds to its targets by recognizing a phosphotyrosine (pY) in a pYxNx peptide motif, requiring an Asn at the +2 position C-terminal to the pY with the residue either side of this Asn being hydrophobic. Structural analysis of the Grb2 SH2 domain in complex with its cognate peptide has shown that the peptide adopts a unique β-turn conformation, unlike the extended conformation that phosphopeptides adopt when bound to other SH2 domains. TrpEF1 (W121) is believed to force the peptide into this unusual conformation conferring this unique specificity to the Grb2 SH2 domain. Using X-ray crystallography, electron paramagnetic resonance (EPR) spectroscopy, and isothermal titration calorimetry (ITC), we describe here a series of experiments that explore the role of TrpEF1 in determining the specificity of the Grb2 SH2 domain. Our results demonstrate that the ligand does not adopt a pre-organized structure before binding to the SH2 domain, rather it is the interaction between the two that imposes the hairpin loop to the peptide. Furthermore, we find that the peptide adopts a similar structure when bound to both the wild-type Grb2 SH2 domain and a TrpEF1Gly mutant. This suggests that TrpEF1 is not the determining factor for the conformation of the phosphopeptide.
- UCL And Birkbeck, Institute of Structural and Molecular Biology, Malet Street, London, WC1E 7HX, United Kingdom.
Organizational Affiliation: