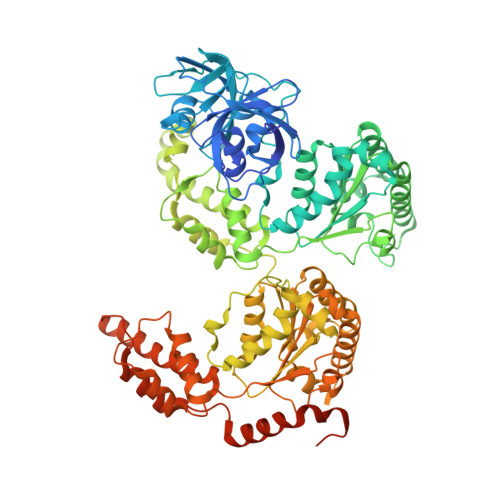
Characterization of an Additional Binding Surface on the p97 N-Terminal Domain Involved in Bipartite Cofactor Interactions.
Hanzelmann, P., Schindelin, H.(2016) Structure 24: 140-147
- PubMed: 26712280 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2015.10.027
- Primary Citation Related Structures:
5C1B - PubMed Abstract:
The type II AAA ATPase p97 interacts with a large number of cofactors that regulate its function by recruiting it to different cellular pathways. Most of the cofactors interact with the N-terminal (N) domain of p97, either via ubiquitin-like domains or short linear binding motifs. While some linear binding motifs form α helices, another group features short stretches of unstructured hydrophobic sequences as found in the so-called SHP (BS1, binding segment 1) motif. Here we present the crystal structure of a SHP-binding motif in complex with p97, which reveals a so far uncharacterized binding site on the p97 N domain that is different from the conserved binding surface of all other known p97 cofactors. This finding explains how cofactors like UFD1/NPL4 and p47 can utilize a bipartite binding mechanism to interact simultaneously with the same p97 monomer via their ubiquitin-like domain and SHP motif.
- Rudolf Virchow Center for Experimental Biomedicine, University of Würzburg, Josef-Schneider-Straße 2, 97080 Würzburg, Germany. Electronic address: petra.haenzelmann@virchow.uni-wuerzburg.de.
Organizational Affiliation: