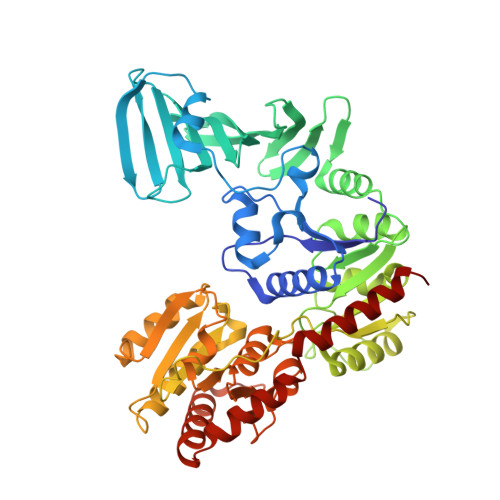
Structure of a Novel O-Linked N-Acetyl-d-glucosamine (O-GlcNAc) Transferase, GtfA, Reveals Insights into the Glycosylation of Pneumococcal Serine-rich Repeat Adhesins.
Shi, W.W., Jiang, Y.L., Zhu, F., Yang, Y.H., Shao, Q.Y., Yang, H.B., Ren, Y.M., Wu, H., Chen, Y., Zhou, C.Z.(2014) J Biological Chem 289: 20898-20907
- PubMed: 24936067 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M114.581934
- Primary Citation Related Structures:
4PQG - PubMed Abstract:
Protein glycosylation catalyzed by the O-GlcNAc transferase (OGT) plays a critical role in various biological processes. In Streptococcus pneumoniae, the core enzyme GtfA and co-activator GtfB form an OGT complex to glycosylate the serine-rich repeat (SRR) of adhesin PsrP (pneumococcal serine-rich repeat protein), which is involved in the infection and pathogenesis. Here we report the 2.0 Å crystal structure of GtfA, revealing a β-meander add-on domain beyond the catalytic domain. It represents a novel add-on domain, which is distinct from the all-α-tetratricopeptide repeats in the only two structure-known OGTs. Structural analyses combined with binding assays indicate that this add-on domain contributes to forming an active GtfA-GtfB complex and recognizing the acceptor protein. In addition, the in vitro glycosylation system enables us to map the O-linkages to the serine residues within the first SRR of PsrP. These findings suggest that fusion with an add-on domain might be a universal mechanism for diverse OGTs that recognize varying acceptor proteins/peptides.