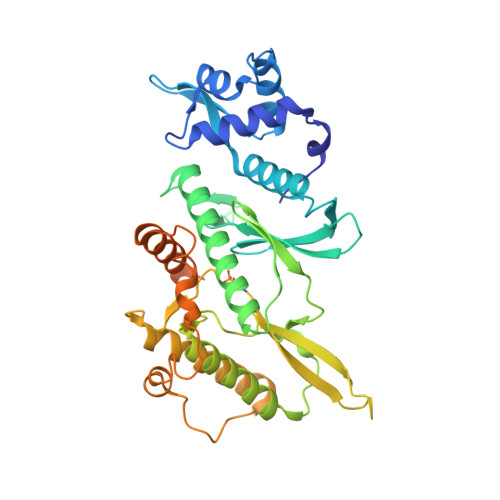
ATPase-dependent role of the atypical kinase Rio2 on the evolving pre-40S ribosomal subunit.
Ferreira-Cerca, S., Sagar, V., Schafer, T., Diop, M., Wesseling, A.M., Lu, H., Chai, E., Hurt, E., Laronde-Leblanc, N.(2012) Nat Struct Mol Biol 19: 1316-1323
- PubMed: 23104056
- DOI: https://doi.org/10.1038/nsmb.2403
- Primary Citation of Related Structures:
4GYG, 4GYI - PubMed Abstract:
Ribosome synthesis involves dynamic association of ribosome-biogenesis factors with evolving preribosomal particles. Rio2 is an atypical protein kinase required for pre-40S subunit maturation. We report the crystal structure of eukaryotic Rio2-ATP-Mg(2+) complex. The active site contains ADP-Mg(2+) and a phosphoaspartate intermediate typically found in Na(+), K(+) and Ca(2+) ATPases but not protein kinases. Consistent with this finding, ctRio2 exhibits a robust ATPase activity in vitro. In vivo, Rio2 docks on the ribosome, with its active site occluded and its flexible loop positioned to interact with the pre-40S subunit. Moreover, Rio2 catalytic activity is required for its dissociation from the ribosome, a necessary step in pre-40S maturation. We propose that phosphoryl transfer from ATP to Asp257 in Rio2's active site and subsequent hydrolysis of the aspartylphosphate could be a trigger to power late cytoplasmic 40S subunit biogenesis.
- Biochemistry Center, University of Heidelberg, Heidelberg, Germany.
Organizational Affiliation: