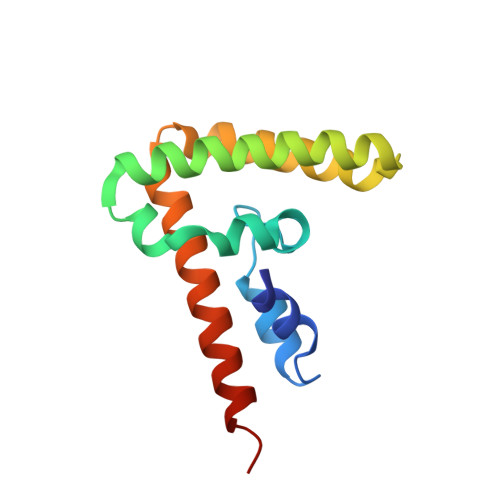
Impact of the N-Terminal Domain of STAT3 in STAT3-Dependent Transcriptional Activity.
Hu, T., Yeh, J.E., Pinello, L., Jacob, J., Chakravarthy, S., Yuan, G.C., Chopra, R., Frank, D.A.(2015) Mol Cell Biol 35: 3284-3300
- PubMed: 26169829 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/MCB.00060-15
- Primary Citation Related Structures:
4ZIA - PubMed Abstract:
The transcription factor STAT3 is constitutively active in many cancers, where it mediates important biological effects, including cell proliferation, differentiation, survival, and angiogenesis. The N-terminal domain (NTD) of STAT3 performs multiple functions, such as cooperative DNA binding, nuclear translocation, and protein-protein interactions. However, it is unclear which subsets of STAT3 target genes depend on the NTD for transcriptional regulation. To identify such genes, we compared gene expression in STAT3-null mouse embryonic fibroblasts (MEFs) stably expressing wild-type STAT3 or STAT3 from which NTD was deleted. NTD deletion reduced the cytokine-induced expression of specific STAT3 target genes by decreasing STAT3 binding to their regulatory regions. To better understand the potential mechanisms of this effect, we determined the crystal structure of the STAT3 NTD and identified a dimer interface responsible for cooperative DNA binding in vitro. We also observed an Ni(2+)-mediated oligomer with an as yet unknown biological function. Mutations on both dimer and Ni(2+)-mediated interfaces affected the cytokine induction of STAT3 target genes. These studies shed light on the role of the NTD in transcriptional regulation by STAT3 and provide a structural template with which to design STAT3 NTD inhibitors with potential therapeutic value.
- Center for Proteomic Chemistry, Novartis Institutes for BioMedical Research, Cambridge, Massachusetts, USA Postdoctoral Program, Novartis Institutes for BioMedical Research, Cambridge, Massachusetts, USA.
Organizational Affiliation: