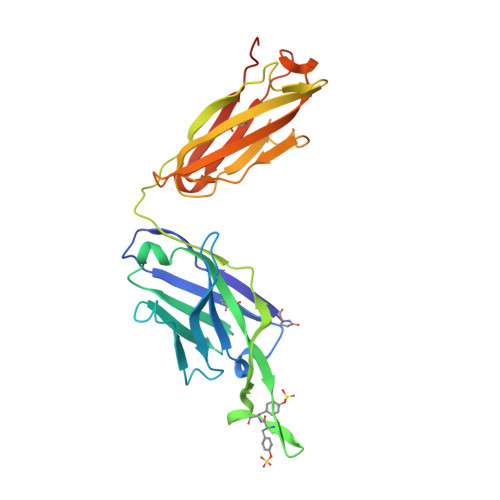
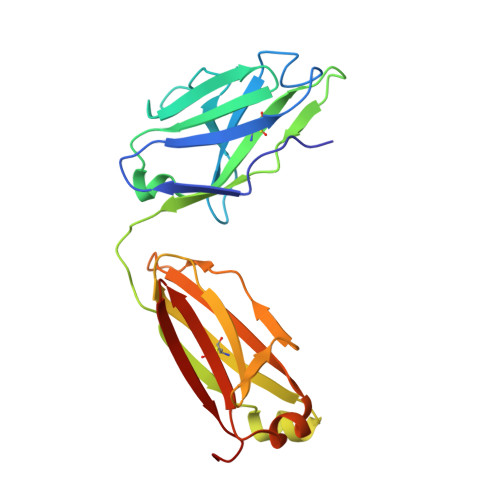
Redesigned HIV antibodies exhibit enhanced neutralizing potency and breadth.
Willis, J.R., Sapparapu, G., Murrell, S., Julien, J.P., Singh, V., King, H.G., Xia, Y., Pickens, J.A., LaBranche, C.C., Slaughter, J.C., Montefiori, D.C., Wilson, I.A., Meiler, J., Crowe, J.E.(2015) J Clin Invest 125: 2523-2531
- PubMed: 25985274 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1172/JCI80693
- Primary Citation Related Structures:
4YAQ - PubMed Abstract:
Several HIV envelope-targeting (Env-targeting) antibodies with broad and potent neutralizing activity have been identified and shown to have unusual features. Of these, the PG9 antibody has a long heavy chain complementarity determining region 3 (HCDR3) and possesses unique structural elements that interact with protein and glycan features of the HIV Env glycoprotein. Here, we used the Rosetta software suite to design variants of the PG9 antibody HCDR3 loop with the goal of identifying variants with increased potency and breadth of neutralization for diverse HIV strains. One variant, designated PG9_N100(F)Y, possessed increased potency and was able to neutralize a diverse set of PG9-resistant HIV strains, including those lacking the Env N160 glycan, which is critical for PG9 binding. An atomic resolution structure of the PG9_N100(F)Y fragment antigen binding (Fab) confirmed that the mutated residue retains the paratope surface when compared with WT PG9. Differential scanning calorimetry experiments revealed that the mutation caused a modest increase in thermodynamic stability of the Fab, a feature predicted by the computational model. Our findings suggest that thermodynamic stabilization of the long HCDR3 in its active conformation is responsible for the increased potency of PG9_N100(F)Y, and strategies aimed at stabilizing this region in other HIV antibodies could become an important approach to in silico optimization of antibodies.