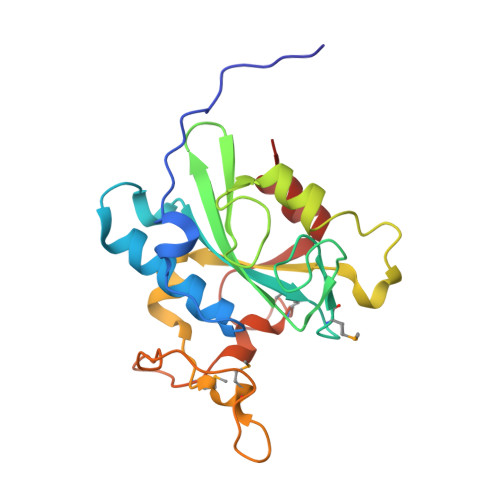
Crystal structure of human protein N-terminal glutamine amidohydrolase, an initial component of the N-end rule pathway.
Park, M.S., Bitto, E., Kim, K.R., Bingman, C.A., Miller, M.D., Kim, H.J., Han, B.W., Phillips, G.N.(2014) PLoS One 9: e111142-e111142
- PubMed: 25356641 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0111142
- Primary Citation Related Structures:
4W79 - PubMed Abstract:
The N-end rule states that half-life of protein is determined by their N-terminal amino acid residue. N-terminal glutamine amidohydrolase (Ntaq) converts N-terminal glutamine to glutamate by eliminating the amine group and plays an essential role in the N-end rule pathway for protein degradation. Here, we report the crystal structure of human Ntaq1 bound with the N-terminus of a symmetry-related Ntaq1 molecule at 1.5 Å resolution. The structure reveals a monomeric globular protein with alpha-beta-alpha three-layer sandwich architecture. The catalytic triad located in the active site, Cys-His-Asp, is highly conserved among Ntaq family and transglutaminases from diverse organisms. The N-terminus of a symmetry-related Ntaq1 molecule bound in the substrate binding cleft and the active site suggest possible substrate binding mode of hNtaq1. Based on our crystal structure of hNtaq1 and docking study with all the tripeptides with N-terminal glutamine, we propose how the peptide backbone recognition patch of hNtaq1 forms nonspecific interactions with N-terminal peptides of substrate proteins. Upon binding of a substrate with N-terminal glutamine, active site catalytic triad mediates the deamination of the N-terminal residue to glutamate by a mechanism analogous to that of cysteine proteases.
- Research Institute of Pharmaceutical Sciences, College of Pharmacy, Seoul National University, Seoul, Korea.
Organizational Affiliation: