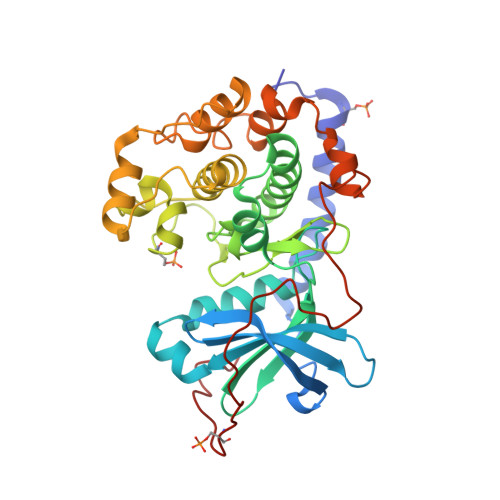
Addressing the Glycine-Rich Loop of Protein Kinases by a Multi-Facetted Interaction Network: Inhibition of Pka and a Pkb Mimic.
Lauber, B.S., Hardegger, L.A., Asraful, A.K., Lund, B.A., Dumele, O., Harder, M., Kuhn, B., Engh, R.A., Diederich, F.(2016) Chemistry 22: 211
- PubMed: 26578105 Search on PubMed
- DOI: https://doi.org/10.1002/chem.201503552
- Primary Citation Related Structures:
4UJ1, 4UJ2, 4UJ9, 4UJA, 4UJB, 4Z83, 4Z84 - PubMed Abstract:
Protein kinases continue to be hot targets in drug discovery research, as they are involved in many essential cellular processes and their deregulation can lead to a variety of diseases. A series of 32 enantiomerically pure inhibitors was synthesized and tested towards protein kinase A (PKA) and protein kinase B mimic PKAB3 (PKA triple mutant). The ligands bind to the hinge region, ribose pocket, and glycine-rich loop at the ATP site. Biological assays showed high potency against PKA, with Ki values in the low nanomolar range. The investigation demonstrates the significance of targeting the often neglected glycine-rich loop for gaining high binding potency. X-ray co-crystal structures revealed a multi-facetted network of ligand-loop interactions for the tightest binders, involving orthogonal dipolar contacts, sulfur and other dispersive contacts, amide-π stacking, and H-bonding to organofluorine, besides efficient water replacement. The network was analyzed in a computational approach.
- Laboratorium für Organische Chemie, ETH Zürich, Vladimir-Prelog-Weg 3, 8093 Zürich (Switzerland), Fax: (+41) 44-632-11-09.
Organizational Affiliation: