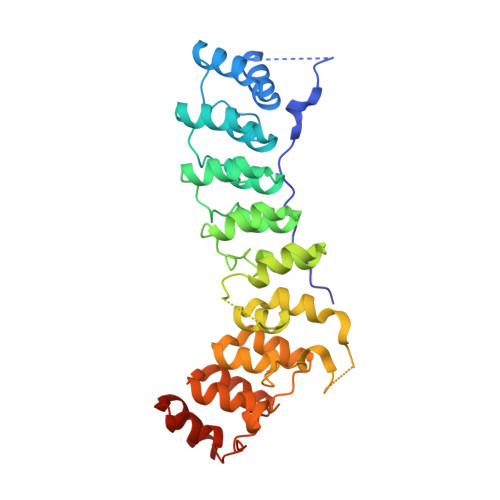
Structural basis of diverse membrane target recognitions by ankyrins.
Wang, C., Wei, Z., Chen, K., Ye, F., Yu, C., Bennett, V., Zhang, M.(2014) Elife 3: e04353
- PubMed: 25383926 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.04353
- Primary Citation Related Structures:
4RLV, 4RLY - PubMed Abstract:
Ankyrin adaptors together with their spectrin partners coordinate diverse ion channels and cell adhesion molecules within plasma membrane domains and thereby promote physiological activities including fast signaling in the heart and nervous system. Ankyrins specifically bind to numerous membrane targets through their 24 ankyrin repeats (ANK repeats), although the mechanism for the facile and independent evolution of these interactions has not been resolved. Here we report the structures of ANK repeats in complex with an inhibitory segment from the C-terminal regulatory domain and with a sodium channel Nav1.2 peptide, respectively, showing that the extended, extremely conserved inner groove spanning the entire ANK repeat solenoid contains multiple target binding sites capable of accommodating target proteins with very diverse sequences via combinatorial usage of these sites. These structures establish a framework for understanding the evolution of ankyrins' membrane targets, with implications for other proteins containing extended ANK repeat domains.
- Division of Life Science, State Key Laboratory of Molecular Neuroscience, Hong Kong University of Science and Technology, Hong Kong, Hong Kong.
Organizational Affiliation: