Interactions between Anticancer trans-Platinum Compounds and Proteins: Crystal Structures and ESI-MS Spectra of Two Protein Adducts of trans-(Dimethylamino)(methylamino)dichloridoplatinum(II).
Messori, L., Marzo, T., Michelucci, E., Russo Krauss, I., Navarro-Ranninger, C., Quiroga, A.G., Merlino, A.(2014) Inorg Chem 53: 7806-7808
- PubMed: 25025479 Search on PubMed
- DOI: https://doi.org/10.1021/ic5012583
- Primary Citation Related Structures:
4QGZ, 4QH3 - PubMed Abstract:
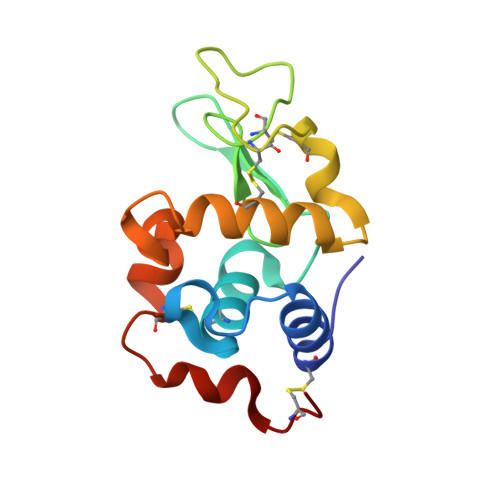
The adducts formed between trans-(dimethylamino)(methylamino)dichloridoplatinum(II), [t-PtCl2(dma)(ma)], and two model proteins, i.e., hen egg white lysozyme and bovine pancreatic ribonuclease, were independently characterized by X-ray crystallography and electrospray ionization mass spectrometry. In these adducts, the Pt(II) center, upon chloride release, coordinates either to histidine or aspartic acid residues while both alkylamino ligands remain bound to the metal. Comparison with the cisplatin derivatives of the same proteins highlights for [t-PtCl2(dma)(ma)] a kind of biomolecular metalation remarkably different from that of cisplatin.
- Department of Chemistry, University of Florence , Via della Lastruccia 3, 50019, Sesto Fiorentino, Italy.
Organizational Affiliation: