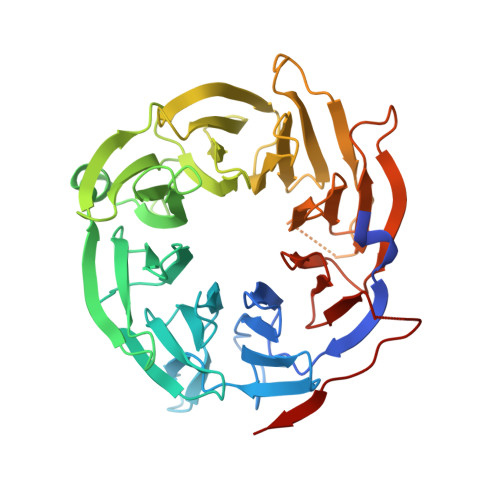
CRL4-like Clr4 complex in Schizosaccharomyces pombe depends on an exposed surface of Dos1 for heterochromatin silencing.
Kuscu, C., Zaratiegui, M., Kim, H.S., Wah, D.A., Martienssen, R.A., Schalch, T., Joshua-Tor, L.(2014) Proc Natl Acad Sci U S A 111: 1795-1800
- PubMed: 24449894 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1313096111
- Primary Citation Related Structures:
4O9D - PubMed Abstract:
Repressive histone H3 lysine 9 methylation (H3K9me) and its recognition by HP1 proteins are necessary for pericentromeric heterochromatin formation. In Schizosaccharomyces pombe, H3K9me deposition depends on the RNAi pathway. Cryptic loci regulator 4 (Clr4), the only known H3K9 methyltransferase in this organism, is a subunit of the Clr4 methyltransferase complex (CLRC), whose composition is reminiscent of a CRL4 type cullin-RING ubiquitin ligase (CRL) including its cullin Cul4, the RING-box protein Pip1, the DNA damage binding protein 1 homolog Rik1, and the DCAF-like protein delocalization of Swi6 1 (Dos1). Dos2 and Stc1 have been proposed to be part of the complex but do not bear similarity to canonical ubiquitin ligase components. CLRC is an active E3 ligase in vitro, and this activity is necessary for heterochromatin assembly in vivo. The similarity between CLRC and the CRLs suggests that the WD repeat protein Dos1 will act to mediate target recognition and substrate specificity for CLRC. Here, we present a pairwise interaction screen that confirms a CRL4-like subunit arrangement and further identifies Dos2 as a central component of the complex and recruiter of Stc1. We determined the crystal structure of the Dos1 WD repeat domain, revealing an eight-bladed β-propeller fold. Functional mapping of the putative target-binding surface of Dos1 identifies key residues required for heterochromatic silencing, consistent with Dos1's role as the specificity factor for the E3 ubiquitin ligase.
- W. M. Keck Structural Biology Laboratory, Cold Spring Harbor, NY 11724.
Organizational Affiliation: