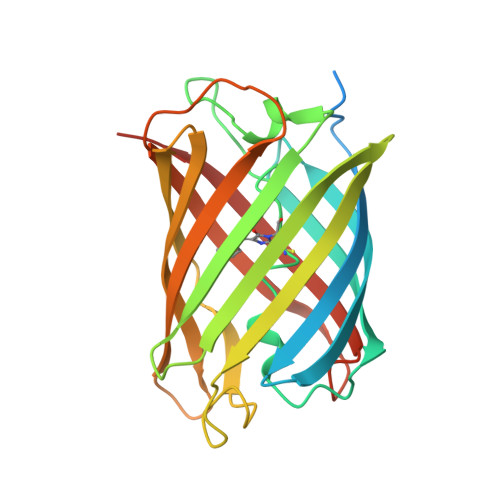
Structural basis for the influence of a single mutation K145N on the oligomerization and photoswitching rate of Dronpa.
Nguyen Bich, N., Moeyaert, B., Van Hecke, K., Dedecker, P., Mizuno, H., Hofkens, J., Van Meervelt, L.(2012) Acta Crystallogr D Biol Crystallogr 68: 1653-1659
- PubMed: 23151630 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444912039686
- Primary Citation Related Structures:
4EMQ - PubMed Abstract:
The crystal structure of the on-state of PDM1-4, a single-mutation variant of the photochromic fluorescent protein Dronpa, is reported at 1.95 Å resolution. PDM1-4 is a Dronpa variant that possesses a slower off-switching rate than Dronpa and thus can effectively increase the image resolution in subdiffraction optical microscopy, although the precise molecular basis for this change has not been elucidated. This work shows that the Lys145Asn mutation in PDM1-4 stabilizes the interface available for dimerization, facilitating oligomerization of the protein. No significant changes were observed in the chromophore environment of PDM1-4 compared with Dronpa, and the ensemble absorption and emission properties of PDM1-4 were highly similar to those of Dronpa. It is proposed that the slower off-switching rate in PDM1-4 is caused by a decrease in the potential flexibility of certain β-strands caused by oligomerization along the AC interface.
- Biomolecular Architecture, Department of Chemistry, KU Leuven, Celestijnenlaan 200F, Leuven (Heverlee), Belgium.
Organizational Affiliation: