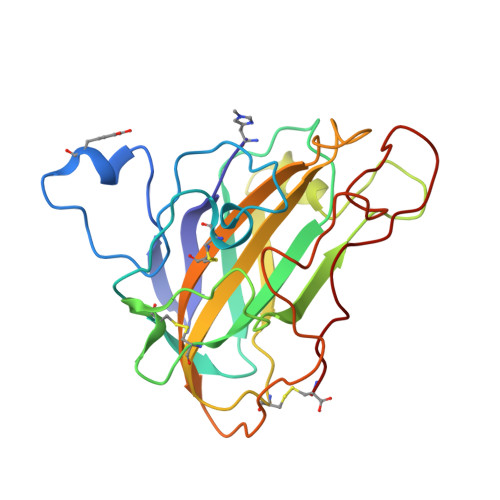
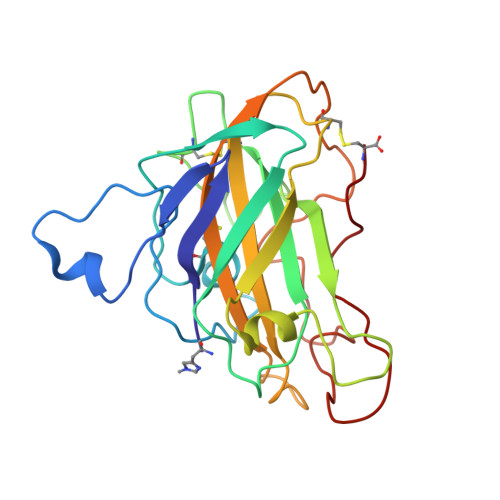
Structural basis for substrate targeting and catalysis by fungal polysaccharide monooxygenases.
Li, X., Beeson, W.T., Phillips, C.M., Marletta, M.A., Cate, J.H.(2012) Structure 20: 1051-1061
- PubMed: 22578542 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2012.04.002
- Primary Citation Related Structures:
4EIR, 4EIS - PubMed Abstract:
The use of cellulases remains a major cost in the production of renewable fuels and chemicals from lignocellulosic biomass. Fungi secrete copper-dependent polysaccharide monooxygenases (PMOs) that oxidatively cleave crystalline cellulose and improve the effectiveness of cellulases. However, the means by which PMOs recognize and cleave their substrates in the plant cell wall remain unclear. Here, we present structures of Neurospora crassa PMO-2 and PMO-3 at 1.10 and 1.37 Å resolution, respectively. In the structures, dioxygen species are found in the active sites, consistent with the proposed cleavage mechanism. Structural and sequence comparisons between PMOs also reveal that the enzyme substrate-binding surfaces contain highly varied aromatic amino acid and glycosylation positions. The structures reported here provide evidence for a wide range of PMO substrate recognition patterns in the plant cell wall, including binding modes that traverse multiple glucan chains.
- California Institute for Quantitative Biosciences, University of California, Berkeley, CA 94720, USA.
Organizational Affiliation: