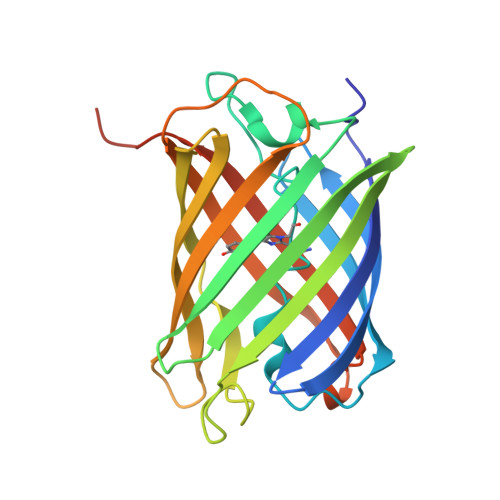
A hinge migration mechanism unlocks the evolution of green-to-red photoconversion in GFP-like proteins.
Kim, H., Zou, T., Modi, C., Dorner, K., Grunkemeyer, T.J., Chen, L., Fromme, R., Matz, M.V., Ozkan, S.B., Wachter, R.M.(2015) Structure 23: 34-43
- PubMed: 25565105 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2014.11.011
- Primary Citation Related Structures:
4DXI, 4DXM, 4DXO, 4DXP - PubMed Abstract:
In proteins, functional divergence involves mutations that modify structure and dynamics. Here we provide experimental evidence for an evolutionary mechanism driven solely by long-range dynamic motions without significant backbone adjustments, catalytic group rearrangements, or changes in subunit assembly. Crystallographic structures were determined for several reconstructed ancestral proteins belonging to a GFP class frequently employed in superresolution microscopy. Their chain flexibility was analyzed using molecular dynamics and perturbation response scanning. The green-to-red photoconvertible phenotype appears to have arisen from a common green ancestor by migration of a knob-like anchoring region away from the active site diagonally across the β barrel fold. The allosterically coupled mutational sites provide active site conformational mobility via epistasis. We propose that light-induced chromophore twisting is enhanced in a reverse-protonated subpopulation, activating internal acid-base chemistry and backbone cleavage to enlarge the chromophore. Dynamics-driven hinge migration may represent a more general platform for the evolution of novel enzyme activities.
- Department of Chemistry and Biochemistry, Arizona State University, Tempe, AZ 85287, USA.
Organizational Affiliation: