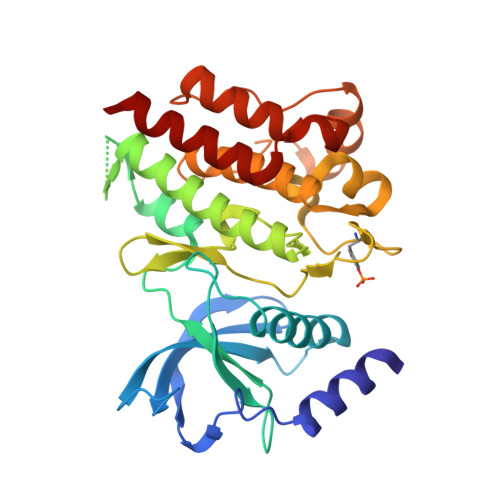
Oncogenic RET kinase domain mutations perturb the autophosphorylation trajectory by enhancing substrate presentation in trans.
Plaza-Menacho, I., Barnouin, K., Goodman, K., Martinez-Torres, R.J., Borg, A., Murray-Rust, J., Mouilleron, S., Knowles, P., McDonald, N.Q.(2014) Mol Cell 53: 738-751
- PubMed: 24560924 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2014.01.015
- Primary Citation Related Structures:
4CKI, 4CKJ - PubMed Abstract:
To decipher the molecular basis for RET kinase activation and oncogenic deregulation, we defined the temporal sequence of RET autophosphorylation by label-free quantitative mass spectrometry. Early autophosphorylation sites map to regions flanking the kinase domain core, while sites within the activation loop only form at later time points. Comparison with oncogenic RET kinase revealed that late autophosphorylation sites become phosphorylated much earlier than wild-type RET, which is due to a combination of an enhanced enzymatic activity, increased ATP affinity, and surprisingly, by providing a better intermolecular substrate. Structural analysis of oncogenic M918T and wild-type RET kinase domains reveal a cis-inhibitory mechanism involving tethering contacts between the glycine-rich loop, activation loop, and αC-helix. Tether mutations only affected substrate presentation but perturbed the autophosphorylation trajectory similar to oncogenic mutations. This study reveals an unappreciated role for oncogenic RET kinase mutations in promoting intermolecular autophosphorylation by enhancing substrate presentation.
- Structural Biology Laboratory, London Research Institute, Cancer Research UK, WC2A 3LY London, UK. Electronic address: ivan.plaza-menacho@cancer.org.uk.
Organizational Affiliation: