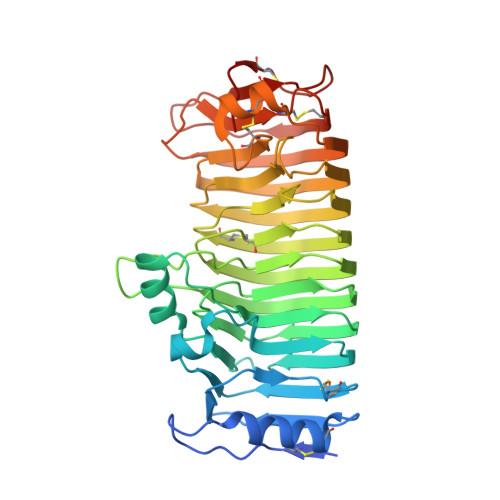
Crystal Structure of Endo-Xylogalacturonan Hydrolase from Aspergillus Tubingensis.
Rozeboom, H.J., Beldman, G., Schols, H.A., Dijkstra, B.W.(2013) FEBS J 280: 6061
- PubMed: 24034788 Search on PubMed
- DOI: https://doi.org/10.1111/febs.12524
- Primary Citation Related Structures:
4C2L - PubMed Abstract:
Endo-xylogalacturonan hydrolase is a member of glycoside hydrolase family 28 (GH28) that hydrolyzes the glycosidic bond between two β-xylose-substituted galacturonic acid residues in pectin. Presented here is the X-ray crystal structure of the endo-xylogalacturonan hydrolase from Aspergillus tubingensis (XghA) at 1.75 Å resolution. The high degree of structural conservation in the active site and catalytic apparatus compared with polygalacturonases indicates that cleavage of the substrate proceeds in essentially the same way as found for the other GH28 enzymes. Molecular modeling of a xylosylated tri-galacturonate in the active site identified the amino acid residues involved in substrate binding. They border a substrate-binding cleft that is much wider than in other polygalacturonases, and can accommodate xylosylated substrates. The most extensive interactions appear to occur at subsite +2, in agreement with the enzyme kinetics results, which showed enhanced activity on substrates with a xylose attached to the galacturonic acid bound at subsite +2. Structural data are available in the Protein Data Bank database under accession number 4C2L.
- Laboratory of Biophysical Chemistry, Groningen Biomolecular Sciences and Biotechnology Institute, University of Groningen, The Netherlands.
Organizational Affiliation: