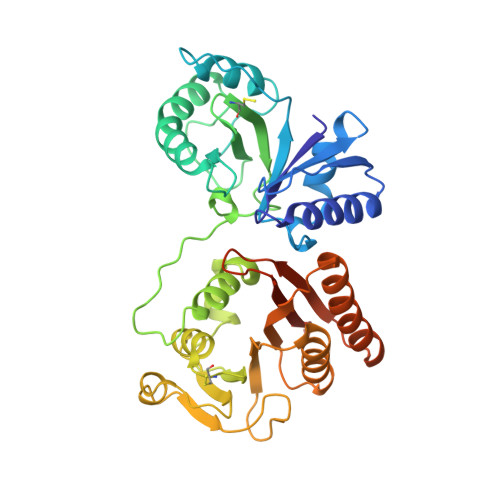
Crystal Structure of Rlmm, the 2'O-Ribose Methyltransferase for C2498 of Escherichia Coli 23S Rrna.
Punekar, A.S., Shepherd, T.R., Liljeruhm, J., Forster, A.C., Selmer, M.(2012) Nucleic Acids Res 40: 10507
- PubMed: 22923526 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gks727
- Primary Citation Related Structures:
4ATN, 4AUK, 4B17 - PubMed Abstract:
RlmM (YgdE) catalyzes the S-adenosyl methionine (AdoMet)-dependent 2'O methylation of C2498 in 23S ribosomal RNA (rRNA) of Escherichia coli. Previous experiments have shown that RlmM is active on 23S rRNA from an RlmM knockout strain but not on mature 50S subunits from the same strain. Here, we demonstrate RlmM methyltransferase (MTase) activity on in vitro transcribed 23S rRNA and its domain V. We have solved crystal structures of E. coli RlmM at 1.9 Å resolution and of an RlmM-AdoMet complex at 2.6 Å resolution. RlmM consists of an N-terminal THUMP domain and a C-terminal catalytic Rossmann-like fold MTase domain in a novel arrangement. The catalytic domain of RlmM is closely related to YiiB, TlyA and fibrillarins, with the second K of the catalytic tetrad KDKE shifted by two residues at the C-terminal end of a beta strand compared with most 2'O MTases. The AdoMet-binding site is open and shallow, suggesting that RNA substrate binding may be required to form a conformation needed for catalysis. A continuous surface of conserved positive charge indicates that RlmM uses one side of the two domains and the inter-domain linker to recognize its RNA substrate.
- Department of Cell and Molecular Biology, Uppsala University, PO Box 596, SE 751 24 Uppsala, Sweden.
Organizational Affiliation: