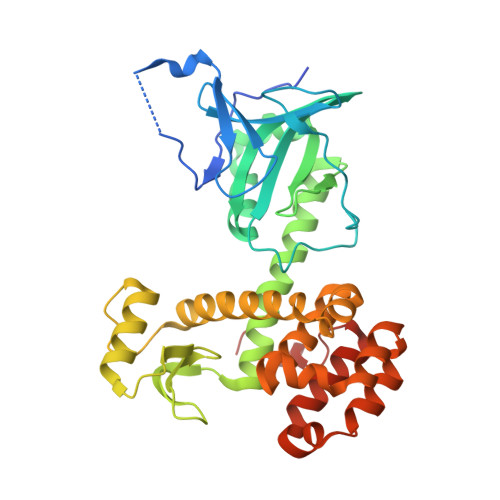
Structure and Mechanistic Studies of Pesticin, a Bacterial Homolog of Phage Lysozymes.
Patzer, S.I., Albrecht, R., Braun, V., Zeth, K.(2012) J Biological Chem 287: 23381
- PubMed: 22593569 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M112.362913
- Primary Citation Related Structures:
4AQN, 4ARJ, 4ARL, 4ARM, 4ARP, 4ARQ - PubMed Abstract:
Yersinia pestis produces and secretes a toxin named pesticin that kills related bacteria of the same niche. Uptake of the bacteriocin is required for activity in the periplasm leading to hydrolysis of peptidoglycan. To understand the uptake mechanism and to investigate the function of pesticin, we combined crystal structures of the wild type enzyme, active site mutants, and a chimera protein with in vivo and in vitro activity assays. Wild type pesticin comprises an elongated N-terminal translocation domain, the intermediate receptor binding domain, and a C-terminal activity domain with structural analogy to lysozyme homologs. The full-length protein is toxic to bacteria when taken up to the target site via the outer or the inner membrane. Uptake studies of deletion mutants in the translocation domain demonstrate their critical size for import. To further test the plasticity of pesticin during uptake into bacterial cells, the activity domain was replaced by T4 lysozyme. Surprisingly, this replacement resulted in an active chimera protein that is not inhibited by the immunity protein Pim. Activity of pesticin and the chimera protein was blocked through introduction of disulfide bonds, which suggests unfolding as the prerequisite to gain access to the periplasm. Pesticin, a muramidase, was characterized by active site mutations demonstrating a similar but not identical residue pattern in comparison with T4 lysozyme.
- Department of Protein Evolution, Max Planck Institute for Developmental Biology, Spemannstrasse 35, 72076 Tübingen, Germany.
Organizational Affiliation: