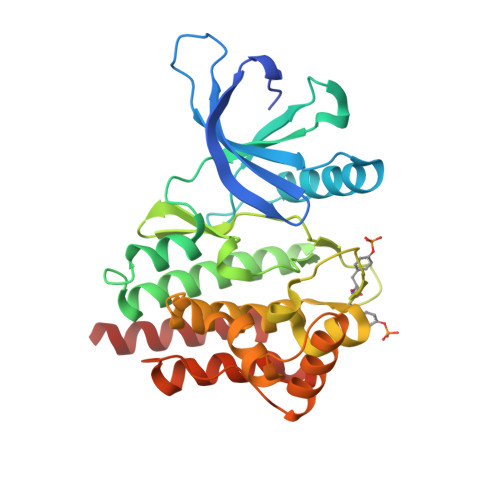
A Selective, Orally Bioavailable 1,2,4-Triazolo[1,5-A]Pyridine-Based Inhibitor of Janus Kinase 2 for Use in Anticancer Therapy: Discovery of Cep-33779.
Dugan, B.J., Gingrich, D.E., Mesaros, E.F., Milkiewicz, K.L., Curry, M.A., Zulli, A.L., Dobrzanski, P., Serdikoff, C., Jan, M., Angeles, T.S., Albom, M.S., Mason, J.L., Aimone, L.D., Meyer, S.L., Huang, Z., Wells-Knecht, K.J., Ator, M.A., Ruggeri, B.A., Dorsey, B.D.(2012) J Med Chem 55: 5243
- PubMed: 22594690 Search on PubMed
- DOI: https://doi.org/10.1021/jm300248q
- Primary Citation Related Structures:
4AQC - PubMed Abstract:
Members of the JAK family of nonreceptor tyrosine kinases play a critical role in the growth and progression of many cancers and in inflammatory diseases. JAK2 has emerged as a leading therapeutic target for oncology, providing a rationale for the development of a selective JAK2 inhibitor. A program to optimize selective JAK2 inhibitors to combat cancer while reducing the risk of immune suppression associated with JAK3 inhibition was undertaken. The structure-activity relationships and biological evaluation of a novel series of compounds based on a 1,2,4-triazolo[1,5-a]pyridine scaffold are reported. Para substitution on the aryl at the C8 position of the core was optimum for JAK2 potency (17). Substitution at the C2 nitrogen position was required for cell potency (21). Interestingly, meta substitution of C2-NH-aryl moiety provided exceptional selectivity for JAK2 over JAK3 (23). These efforts led to the discovery of CEP-33779 (29), a novel, selective, and orally bioavailable inhibitor of JAK2.
- Worldwide Discovery Research, Cephalon, Inc., 145 Brandywine Parkway, West Chester, Pennsylvania 19380, United States. bdugan@cephalon.com
Organizational Affiliation: