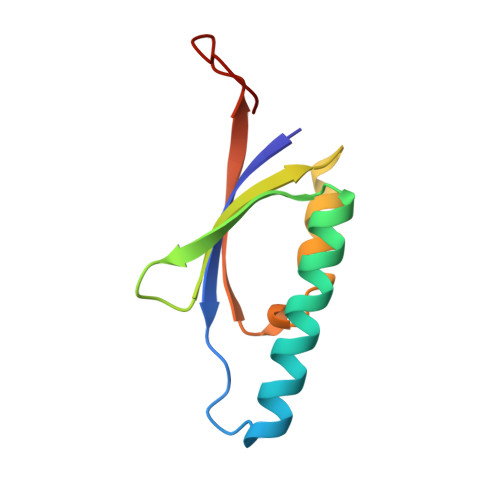
Crystal Structure and Catalytic Mechanism of Chloromuconolactone Dehalogenase Clcf from Rhodococcus Opacus 1Cp.
Roth, C., Janosch, A.D., Kaschabek, S.R., Schloemann, M., Straeter, N.(2013) Mol Microbiol 88: 254
- PubMed: 23421784 Search on PubMed
- DOI: https://doi.org/10.1111/mmi.12182
- Primary Citation Related Structures:
3ZNJ, 3ZNU, 3ZO7 - PubMed Abstract:
The actinobacterium Rhodococcus opacus 1CP possesses a so far unique variant of the modified 3-oxoadipate pathway for 3-chlorocatechol degradation. One important feature is the novel dehalogenase ClcF, which converts (4R,5S)-5-chloromuconolactone to E-dienelactone. ClcF is related to muconolactone isomerase (MLI, EC 5.3.3.4). The enzyme has a ferredoxin-type fold and forms a homodecamer of 52-symmetry, typical for the MLI family. The active site is formed by residues from two monomers. The complex structure of an E27A variant with bound substrate in conjunction with mutational studies indicate that E27 acts as the proton acceptor in a univalent single-base syn-dehydrohalogenation mechanism. Despite the evolutionary specialization of ClcF, the conserved active-site structures suggest that the proposed mechanism is representative for the MLI family. Furthermore, ClcF represents a novel type of dehalogenase based on an isomerase scaffold.
- Center for Biotechnology and Biomedicine, Institute of Bioanalytical Chemistry, Faculty of Chemistry and Mineralogy, University of Leipzig, Deutscher Platz 5, 04103 Leipzig, Germany.
Organizational Affiliation: