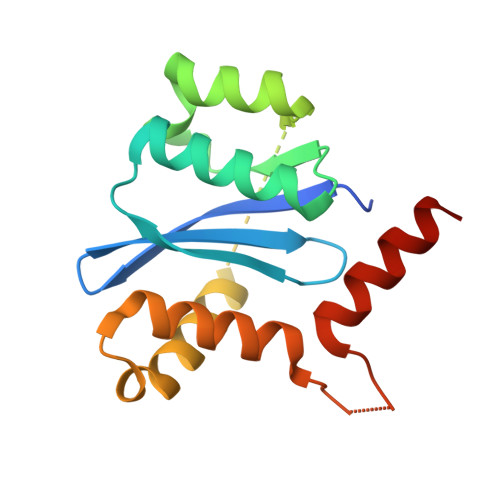
Fragment-based design of ligands targeting a novel site on the integrase enzyme of human immunodeficiency virus 1
Wielens, J., Headey, S.J., Deadman, J.J., Rhodes, D.I., Parker, M.W., Chalmers, D.K., Scanlon, M.J.(2011) ChemMedChem 6: 258-261
- PubMed: 21275048
- DOI: https://doi.org/10.1002/cmdc.201000483
- Primary Citation of Related Structures:
3AO1, 3AO2, 3AO3, 3AO4, 3AO5, 3OVN - Medicinal Chemistry and Drug Action, Monash Institute of Pharmaceutical Sciences, Monash University, 381 Royal Parade, Parkville 3052, Australia.
Organizational Affiliation: