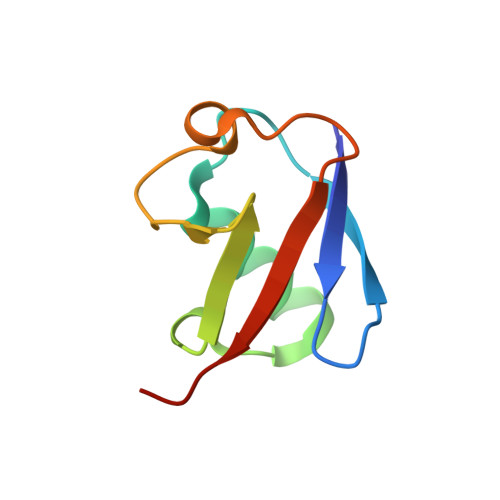
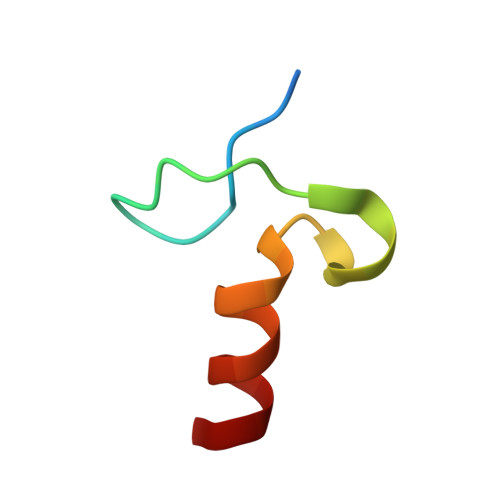
Specific recognition of linear polyubiquitin by A20 zinc finger 7 is involved in NF-kappaB regulation
Tokunaga, F., Nishimasu, H., Ishitani, R., Goto, E., Noguchi, T., Mio, K., Kamei, K., Ma, A., Iwai, K., Nureki, O.(2012) EMBO J 31: 3856-3870
- PubMed: 23032187 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/emboj.2012.241
- Primary Citation Related Structures:
3VUW, 3VUX, 3VUY - PubMed Abstract:
LUBAC (linear ubiquitin chain assembly complex) activates the canonical NF-κB pathway through linear polyubiquitination of NEMO (NF-κB essential modulator, also known as IKKγ) and RIP1. However, the regulatory mechanism of LUBAC-mediated NF-κB activation remains elusive. Here, we show that A20 suppresses LUBAC-mediated NF-κB activation by binding linear polyubiquitin via the C-terminal seventh zinc finger (ZF7), whereas CYLD suppresses it through deubiquitinase (DUB) activity. We determined the crystal structures of A20 ZF7 in complex with linear diubiquitin at 1.70-1.98 Å resolutions. The crystal structures revealed that A20 ZF7 simultaneously recognizes the Met1-linked proximal and distal ubiquitins, and that genetic mutations associated with B cell lymphomas map to the ubiquitin-binding sites. Our functional analysis indicated that the binding of A20 ZF7 to linear polyubiquitin contributes to the recruitment of A20 into a TNF receptor (TNFR) signalling complex containing LUBAC and IκB kinase (IKK), which results in NF-κB suppression. These findings provide new insight into the regulation of immune and inflammatory responses.
- Laboratory of Molecular Cell Biology, Institute for Molecular and Cellular Regulation, Gunma University, Maebashi, Gunma, Japan. ftokunaga@gunma-u.ac.jp
Organizational Affiliation: