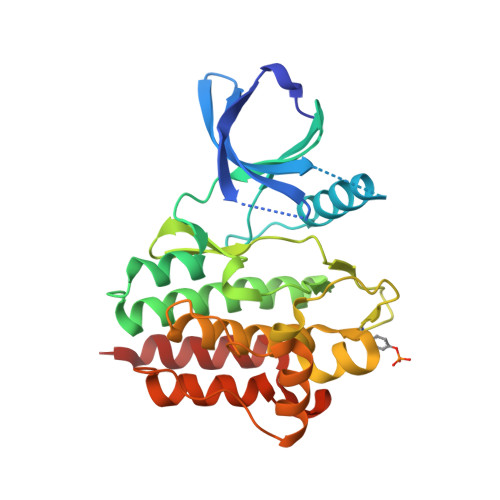
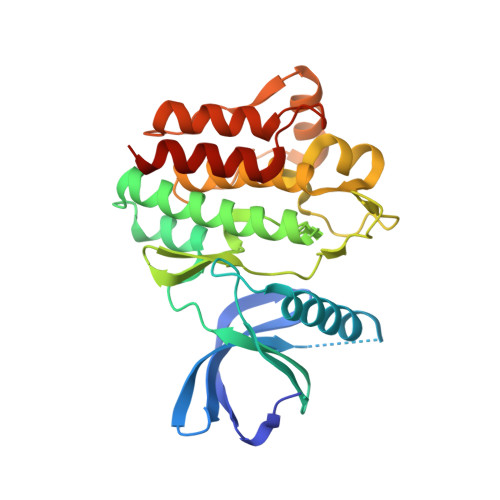
Discovery of GSK143, a highly potent, selective and orally efficacious spleen tyrosine kinase inhibitor.
Liddle, J., Atkinson, F.L., Barker, M.D., Carter, P.S., Curtis, N.R., Davis, R.P., Douault, C., Dickson, M.C., Elwes, D., Garton, N.S., Gray, M., Hayhow, T.G., Hobbs, C.I., Jones, E., Leach, S., Leavens, K., Lewis, H.D., McCleary, S., Neu, M., Patel, V.K., Preston, A.G., Ramirez-Molina, C., Shipley, T.J., Skone, P.A., Smithers, N., Somers, D.O., Walker, A.L., Watson, R.J., Weingarten, G.G.(2011) Bioorg Med Chem Lett 21: 6188-6194
- PubMed: 21903390 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2011.07.082
- Primary Citation Related Structures:
3SRV, 4YJO, 4YJP, 4YJQ, 4YJR, 4YJS, 4YJT, 4YJU - PubMed Abstract:
The lead optimisation of the diaminopyrimidine carboxamide series of spleen tyrosine kinase inhibitors is described. The medicinal chemistry strategy was focused on optimising the human whole blood activity whilst achieving a sufficient margin over liability kinases and hERG activity. GSK143 is a potent and highly selective SYK inhibitor showing good efficacy in the rat Arthus model.
- GlaxoSmithKline R&D, Medicines Research Centre, Gunnels Wood Road, Stevenage, Hertfordshire, UK. john.2.liddle@gsk.com
Organizational Affiliation: