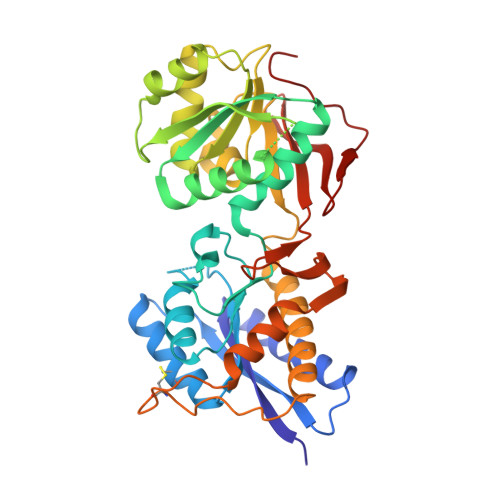
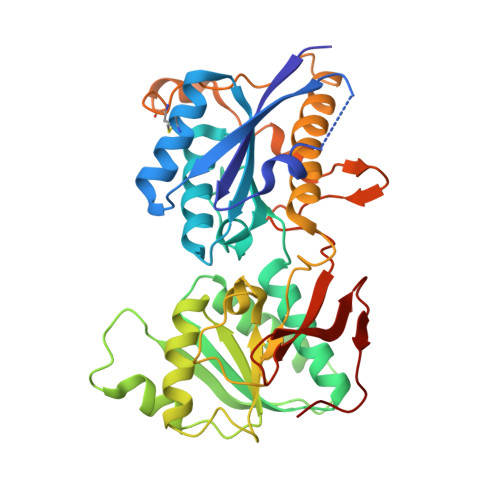
Subunit arrangement and phenylethanolamine binding in GluN1/GluN2B NMDA receptors.
Karakas, E., Simorowski, N., Furukawa, H.(2011) Nature 475: 249-253
- PubMed: 21677647 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature10180
- Primary Citation Related Structures:
3QEK, 3QEL, 3QEM - PubMed Abstract:
Since it was discovered that the anti-hypertensive agent ifenprodil has neuroprotective activity through its effects on NMDA (N-methyl-D-aspartate) receptors, a determined effort has been made to understand the mechanism of action and to develop improved therapeutic compounds on the basis of this knowledge. Neurotransmission mediated by NMDA receptors is essential for basic brain development and function. These receptors form heteromeric ion channels and become activated after concurrent binding of glycine and glutamate to the GluN1 and GluN2 subunits, respectively. A functional hallmark of NMDA receptors is that their ion-channel activity is allosterically regulated by binding of small compounds to the amino-terminal domain (ATD) in a subtype-specific manner. Ifenprodil and related phenylethanolamine compounds, which specifically inhibit GluN1 and GluN2B NMDA receptors, have been intensely studied for their potential use in the treatment of various neurological disorders and diseases, including depression, Alzheimer's disease and Parkinson's disease. Despite considerable enthusiasm, mechanisms underlying the recognition of phenylethanolamines and ATD-mediated allosteric inhibition remain limited owing to a lack of structural information. Here we report that the GluN1 and GluN2B ATDs form a heterodimer and that phenylethanolamine binds at the interface between GluN1 and GluN2B, rather than within the GluN2B cleft. The crystal structure of the heterodimer formed between the GluN1b ATD from Xenopus laevis and the GluN2B ATD from Rattus norvegicus shows a highly distinct pattern of subunit arrangement that is different from the arrangements observed in homodimeric non-NMDA receptors and reveals the molecular determinants for phenylethanolamine binding. Restriction of domain movement in the bi-lobed structure of the GluN2B ATD, by engineering of an inter-subunit disulphide bond, markedly decreases sensitivity to ifenprodil, indicating that conformational freedom in the GluN2B ATD is essential for ifenprodil-mediated allosteric inhibition of NMDA receptors. These findings pave the way for improving the design of subtype-specific compounds with therapeutic value for neurological disorders and diseases.
- Cold Spring Harbor Laboratory, WM Keck Structural Biology Laboratory, 1 Bungtown Road, Cold Spring Harbor, New York 11724, USA.
Organizational Affiliation: