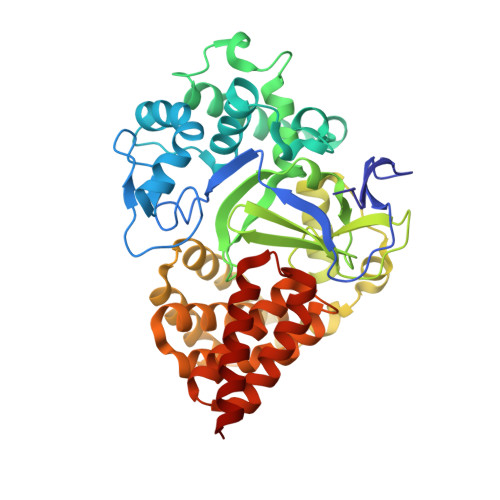
Structural Insights into the Autoinhibition and Posttranslational Activation of Histone Methyltransferase SmyD3.
Sirinupong, N., Brunzelle, J., Doko, E., Yang, Z.(2011) J Mol Biology 406: 149-159
- PubMed: 21167177 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2010.12.014
- Primary Citation Related Structures:
3PDN - PubMed Abstract:
The SmyD family represents a new class of chromatin regulators that is important in heart and skeletal muscle development. However, the critical questions regarding how they are regulated posttranslationally remain largely unknown. We previously suggested that the histone methyltransferase activity of SmyD1, a vital myogenic regulator, appears to be regulated by autoinhibition and that the possible hinge motion of the conserved C-terminal domain (CTD) might be central to the maintenance and release of the autoinhibition. However, the lack of direct evidence of the hinge motion has limited our further understanding of this autoinhibitory mechanism. Here, we report the crystal structure of full-length SmyD3 in complex with the methyltransferase inhibitor sinefungin at 1.7 Å. SmyD3 has a two-lobed structure with the substrate binding cleft located at the bottom of a 15-Å-deep crevice formed between the N- and C-terminal lobes. Comparison of SmyD3 and SmyD1 clearly suggests that the CTD can undergo a large hinge-bending motion that defines two distinct conformations: SmyD3 adopts a closed conformation with the CTD partially blocking the substrate binding cleft; in contrast, SmyD1 appears to represent an open form, where the CTD swings out by ∼12 Å from the N-terminal lobe, forming an open cleft with the active site completely exposed. Overall, these findings provide novel structural insights into the mechanism that modulates the activity of the SmyD proteins and support the observation that a posttranslational activation, such as by molecular chaperon Hsp90, is required to potentiate the proteins.
- Department of Biochemistry and Molecular Biology, Wayne State University School of Medicine, Detroit, MI 48201, USA.
Organizational Affiliation: