Structural basis of thrombin-mediated factor V activation: the Glu666-Glu672 sequence is critical for processing at the heavy chain-B domain junction.
Corral-Rodriguez, M.A., Bock, P.E., Hernandez-Carvajal, E., Gutierrez-Gallego, R., Fuentes-Prior, P.(2011) Blood 117: 7164-7173
- PubMed: 21555742 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1182/blood-2010-10-315309
- Primary Citation Related Structures:
3P6Z, 3P70 - PubMed Abstract:
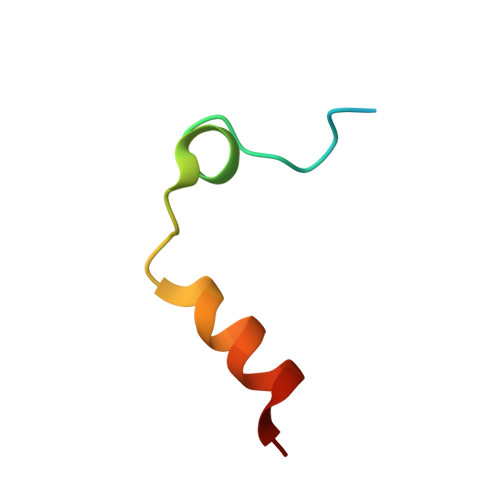
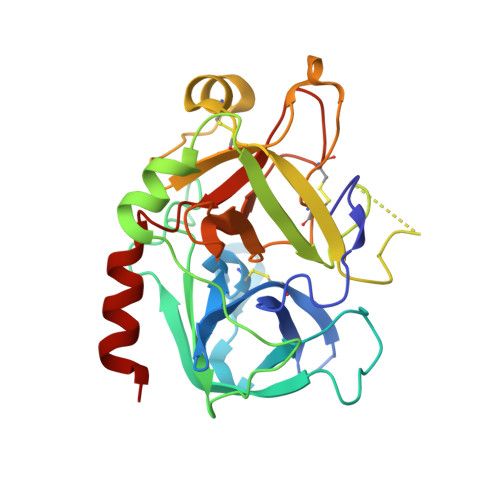
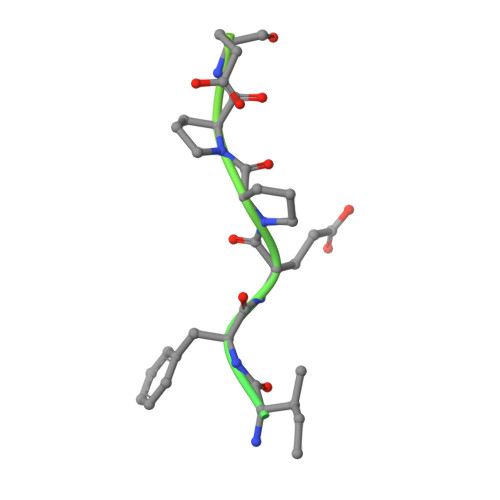
Thrombin-catalyzed activation of coagulation factor V (FV) is an essential positive feedback reaction within the blood clotting system. Efficient processing at the N- (Arg(709)-Ser(710)) and C-terminal activation cleavage sites (Arg(1545)-Ser(1546)) requires initial substrate interactions with 2 clusters of positively charged residues on the proteinase surface, exosites I and II. We addressed the mechanism of activation of human factor V (FV) using peptides that cover the entire acidic regions preceding these cleavage sites, FV (657-709)/ (FVa2) and FV(1481-1545)/(FVa3). FVa2 appears to interact mostly with exosite I, while both exosites are involved in interactions with the C-terminal linker. The 1.7-Å crystal structure of irreversibly inhibited thrombin bound to FVa2 unambiguously reveals docking of FV residues Glu(666)-Glu(672) to exosite I. These findings were confirmed in a second, medium-resolution structure of FVa2 bound to the benzamidine-inhibited proteinase. Our results suggest that the acidic A2-B domain linker is involved in major interactions with thrombin during cofactor activation, with its more N-terminal hirudin-like sequence playing a critical role. Modeling experiments indicate that FVa2, and likely also FVa3, wrap around thrombin in productive thrombin·FV complexes that cover a large surface of the activator to engage the active site.
- Institute for Biomedical Research, Hospital de la Santa Creu i Sant Pau, Barcelona, Spain.
Organizational Affiliation: