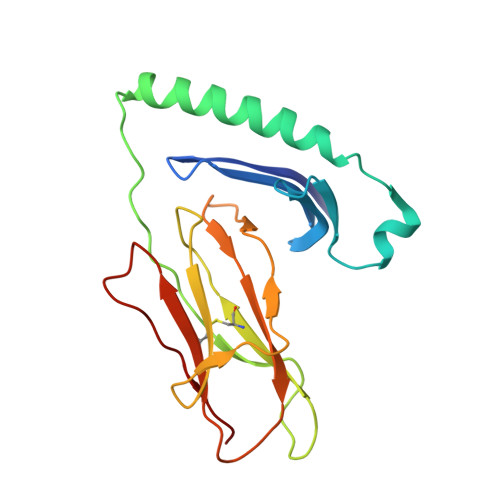
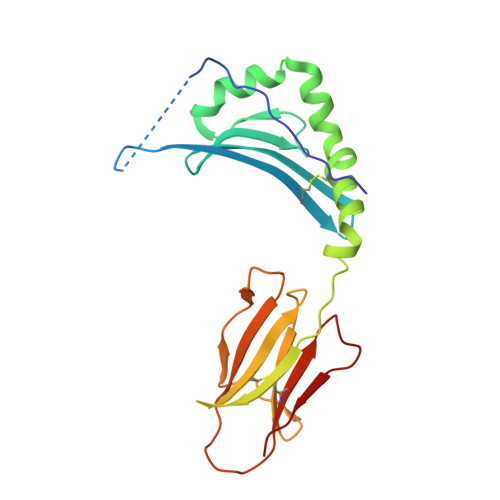
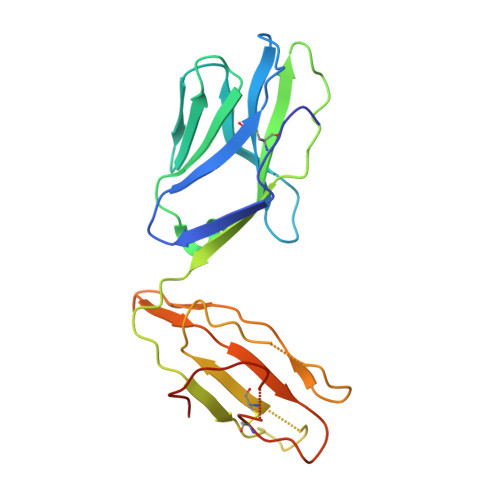
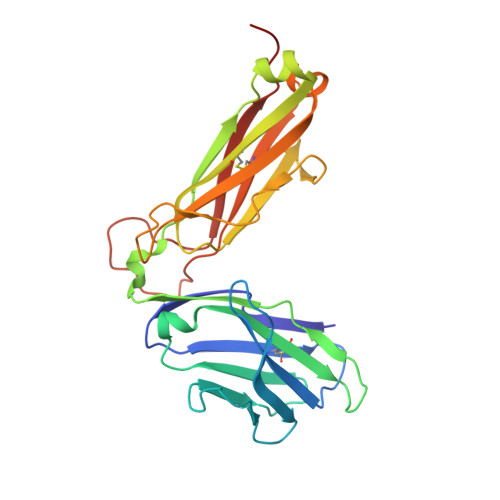
Structure of a TCR with high affinity for self-antigen reveals basis for escape from negative selection.
Yin, Y., Li, Y., Kerzic, M.C., Martin, R., Mariuzza, R.A.(2011) EMBO J 30: 1137-1148
- PubMed: 21297580 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/emboj.2011.21
- Primary Citation Related Structures:
3O6F - PubMed Abstract:
The failure to eliminate self-reactive T cells during negative selection is a prerequisite for autoimmunity. To escape deletion, autoreactive T-cell receptors (TCRs) may form unstable complexes with self-peptide-MHC by adopting suboptimal binding topologies compared with anti-microbial TCRs. Alternatively, escape can occur by weak binding between self-peptides and MHC. We determined the structure of a human autoimmune TCR (MS2-3C8) bound to a self-peptide from myelin basic protein (MBP) and the multiple sclerosis-associated MHC molecule HLA-DR4. MBP is loosely accommodated in the HLA-DR4-binding groove, accounting for its low affinity. Conversely, MS2-3C8 binds MBP-DR4 as tightly as the most avid anti-microbial TCRs. MS2-3C8 engages self-antigen via a docking mode that resembles the optimal topology of anti-foreign TCRs, but is distinct from that of other autoreactive TCRs. Combined with a unique CDR3β conformation, this docking mode compensates for the weak binding of MBP to HLA-DR4 by maximizing interactions between MS2-3C8 and MBP. Thus, the MS2-3C8-MBP-DR4 complex reveals the basis for an alternative strategy whereby autoreactive T cells escape negative selection, yet retain the ability to initiate autoimmunity.
- Institute for Bioscience and Biotechnology Research, University of Maryland, WM Keck Laboratory for Structural Biology, Rockville, MD, USA.
Organizational Affiliation: