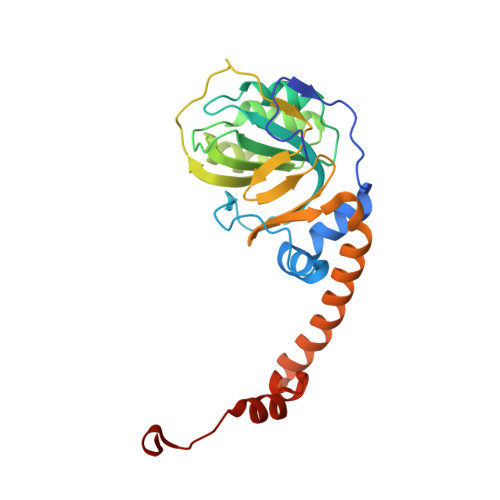
Structural and Kinetic Characterization of 4-Hydroxy-4-methyl-2-oxoglutarate/4-Carboxy-4-hydroxy-2-oxoadipate Aldolase, a Protocatechuate Degradation Enzyme Evolutionarily Convergent with the HpaI and DmpG Pyruvate Aldolases.
Wang, W., Mazurkewich, S., Kimber, M.S., Seah, S.Y.(2010) J Biological Chem 285: 36608-36615
- PubMed: 20843800 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.159509
- Primary Citation Related Structures:
3NOJ - PubMed Abstract:
4-Hydroxy-4-methyl-2-oxoglutarate/4-carboxy-4-hydroxy-2-oxoadipate (HMG/CHA) aldolase from Pseudomonas putida F1 catalyzes the last step of the bacterial protocatechuate 4,5-cleavage pathway. The preferred substrates of the enzyme are 2-keto-4-hydroxy acids with a 4-carboxylate substitution. The enzyme also exhibits oxaloacetate decarboxylation and pyruvate α-proton exchange activity. Sodium oxalate is a competitive inhibitor of the aldolase reaction. The pH dependence of k(cat)/K(m) and k(cat) for the enzyme is consistent with a single deprotonation with pK(a) values of 8.0 ± 0.1 and 7.0 ± 0.1 for free enzyme and enzyme substrate complex, respectively. The 1.8 Å x-ray structure shows a four-layered α-β-β-α sandwich structure with the active site at the interface of two adjacent subunits of a hexamer; this fold resembles the RNase E inhibitor, RraA, but is novel for an aldolase. The catalytic site contains a magnesium ion ligated by Asp-124 as well as three water molecules bound by Asp-102 and Glu-199'. A pyruvate molecule binds the magnesium ion through both carboxylate and keto oxygen atoms, completing the octahedral geometry. The carbonyl oxygen also forms hydrogen bonds with the guanadinium group of Arg-123, which site-directed mutagenesis confirms is essential for catalysis. A mechanism for HMG/CHA aldolase is proposed on the basis of the structure, kinetics, and previously established features of other aldolase mechanisms.
- Department of Molecular and Cellular Biology, University of Guelph, Guelph, Ontario N1G 2W1, Canada.
Organizational Affiliation: