Crystal Structure of Monomeric Kusabira-Orange (MKO), Orange-Emitting GFP-like Protein, at pH 7.5
Ebisawa, T., Yamamura, A., Ohtsuka, J., Kameda, Y., Hayakawa, K., Nagata, K., Tanokura, M.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Fluorescent protein | 218 | Lithophyllon concinna | Mutation(s): 1 Gene Names: mKO | 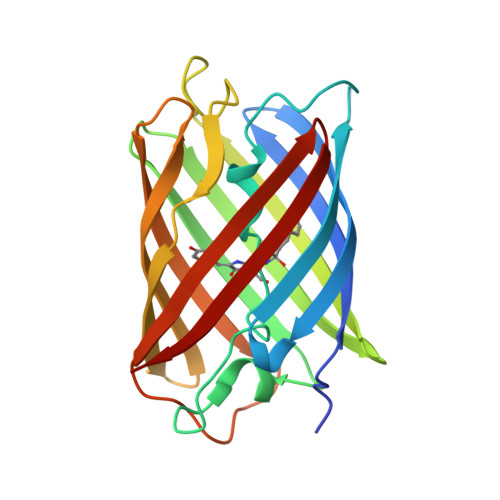 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q6I7B2 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 67.52 | α = 90 |
| b = 83.76 | β = 110.75 |
| c = 82.18 | γ = 90 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| PDB_EXTRACT | data extraction |
| ADSC | data collection |
| XDS | data reduction |
| XDS | data scaling |
| MOLREP | phasing |