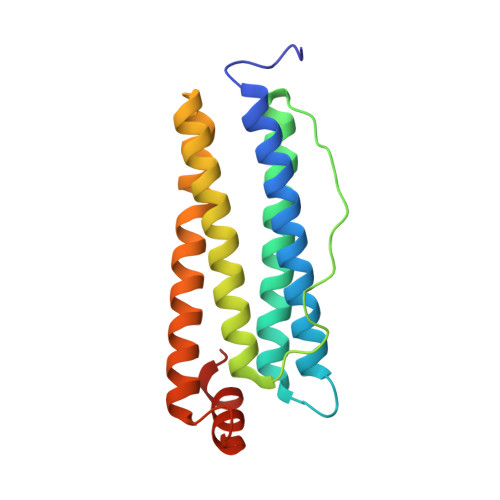
Moving Metal Ions through Ferritin-Protein Nanocages from Three-Fold Pores to Catalytic Sites.
Tosha, T., Ng, H.L., Bhattasali, O., Alber, T., Theil, E.C.(2010) J Am Chem Soc 132: 14562-14569
- PubMed: 20866049 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/ja105583d
- Primary Citation Related Structures:
3KA3, 3KA4, 3KA6, 3KA8 - PubMed Abstract:
Ferritin nanocages synthesize ferric oxide minerals, containing hundreds to thousands of Fe(III) diferric oxo/hydroxo complexes, by reactions of Fe(II) ions with O(2) at multiple di-iron catalytic centers. Ferric-oxy multimers, tetramers, and/or larger mineral nuclei form during postcatalytic transit through the protein cage, and mineral accretion occurs in the central cavity. We determined how Fe(II) substrates can access catalytic sites using frog M ferritins, active and inactivated by ligand substitution, crystallized with 2.0 M Mg(II) ± 0.1 M Co(II) for Co(II)-selective sites. Co(II) inhibited Fe(II) oxidation. High-resolution (<1.5 Å) crystal structures show (1) a line of metal ions, 15 Å long, which penetrates the cage and defines ion channels and internal pores to the nanocavity that link external pores to the cage interior, (2) metal ions near negatively charged residues at the channel exits and along the inner cavity surface that model Fe(II) transit to active sites, and (3) alternate side-chain conformations, absent in ferritins with catalysis eliminated by amino acid substitution, which support current models of protein dynamics and explain changes in Fe-Fe distances observed during catalysis. The new structural data identify a ∼27-Å path Fe(II) ions can follow through ferritin entry channels between external pores and the central cavity and along the cavity surface to the active sites where mineral synthesis begins. This "bucket brigade" for Fe(II) ion access to the ferritin catalytic sites not only increases understanding of biological nanomineral synthesis but also reveals unexpected design principles for protein cage-based catalysts and nanomaterials.
- Council on Bioiron at Children's Hospital Oakland Research Institute, 5700 Martin Luther King Jr. Way, Oakland, California 94609, USA.
Organizational Affiliation: