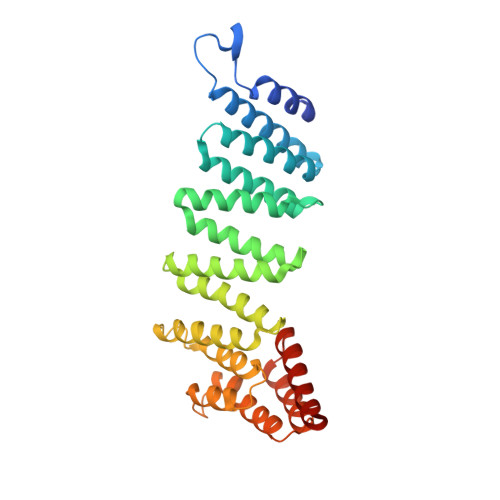
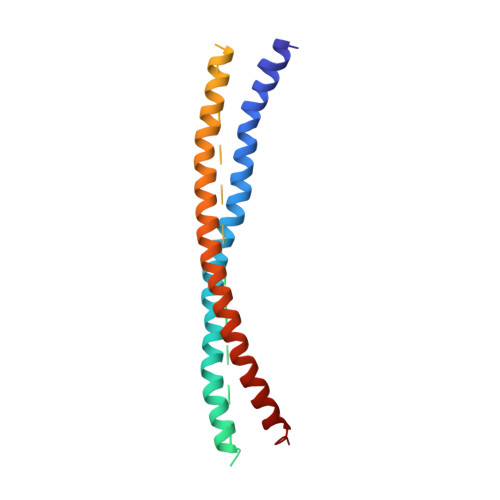
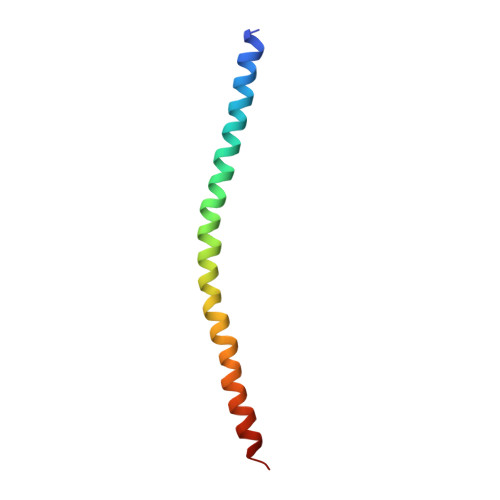Mechanistic insights into the recycling machine of the SNARE complex.
Zhao, M., Wu, S., Zhou, Q., Vivona, S., Cipriano, D.J., Cheng, Y., Brunger, A.T.(2015) Nature 518: 61-67
- PubMed: 25581794 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature14148
- Primary Citation Related Structures:
3J94, 3J95, 3J96, 3J97, 3J98, 3J99 - PubMed Abstract:
Evolutionarily conserved SNARE (soluble N-ethylmaleimide sensitive factor attachment protein receptors) proteins form a complex that drives membrane fusion in eukaryotes. The ATPase NSF (N-ethylmaleimide sensitive factor), together with SNAPs (soluble NSF attachment protein), disassembles the SNARE complex into its protein components, making individual SNAREs available for subsequent rounds of fusion. Here we report structures of ATP- and ADP-bound NSF, and the NSF/SNAP/SNARE (20S) supercomplex determined by single-particle electron cryomicroscopy at near-atomic to sub-nanometre resolution without imposing symmetry. Large, potentially force-generating, conformational differences exist between ATP- and ADP-bound NSF. The 20S supercomplex exhibits broken symmetry, transitioning from six-fold symmetry of the NSF ATPase domains to pseudo four-fold symmetry of the SNARE complex. SNAPs interact with the SNARE complex with an opposite structural twist, suggesting an unwinding mechanism. The interfaces between NSF, SNAPs, and SNAREs exhibit characteristic electrostatic patterns, suggesting how one NSF/SNAP species can act on many different SNARE complexes.
- Department of Molecular and Cellular Physiology, Howard Hughes Medical Institute, Stanford University, Stanford, California 94305, USA.
Organizational Affiliation: