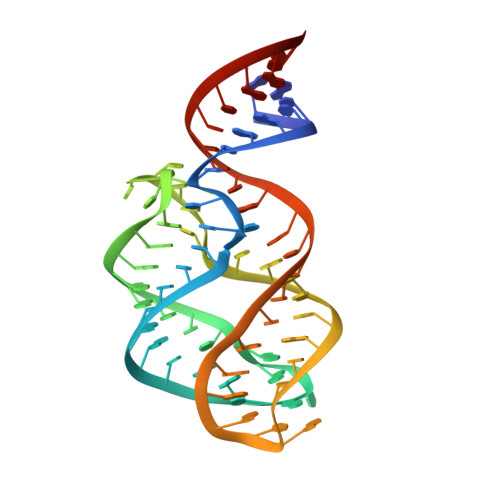
Riboswitch structure: an internal residue mimicking the purine ligand.
Delfosse, V., Bouchard, P., Bonneau, E., Dagenais, P., Lemay, J.F., Lafontaine, D.A., Legault, P.(2010) Nucleic Acids Res 38: 2057-2068
- PubMed: 20022916 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkp1080
- Primary Citation Related Structures:
3IVN - PubMed Abstract:
The adenine and guanine riboswitches regulate gene expression in response to their purine ligand. X-ray structures of the aptamer moiety of these riboswitches are characterized by a compact fold in which the ligand forms a Watson-Crick base pair with residue 65. Phylogenetic analyses revealed a strict restriction at position 39 of the aptamer that prevents the G39-C65 and A39-U65 combinations, and mutational studies indicate that aptamers with these sequence combinations are impaired for ligand binding. In order to investigate the rationale for sequence conservation at residue 39, structural characterization of the U65C mutant from Bacillus subtilis pbuE adenine riboswitch aptamer was undertaken. NMR spectroscopy and X-ray crystallography studies demonstrate that the U65C mutant adopts a compact ligand-free structure, in which G39 occupies the ligand-binding site of purine riboswitch aptamers. These studies present a remarkable example of a mutant RNA aptamer that adopts a native-like fold by means of ligand mimicking and explain why this mutant is impaired for ligand binding. Furthermore, this work provides a specific insight into how the natural sequence has evolved through selection of nucleotide identities that contribute to formation of the ligand-bound state, but ensures that the ligand-free state remains in an active conformation.
- Département de Biochimie, Université de Montréal, C.P. 6128, Succursale Centre-Ville, Montréal, Québec, H3C 3J7, Canada.
Organizational Affiliation: