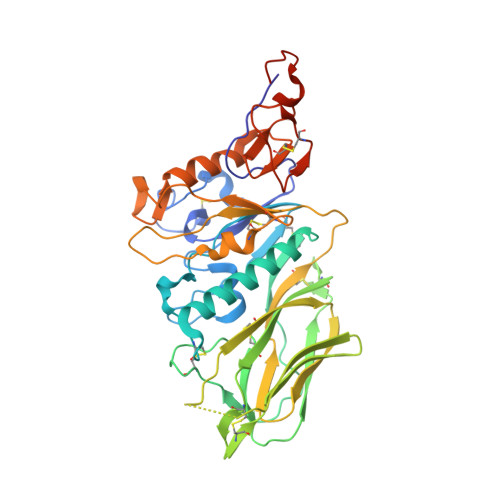
Structural basis for ligand and substrate recognition by torovirus hemagglutinin esterases
Langereis, M.A., Zeng, Q.H., Gerwig, G.J., Frey, B., von Itzstein, M., Kamerling, J.P., de Groot, R.J., Huizinga, E.G.(2009) Proc Natl Acad Sci U S A 106: 15897-15902
- PubMed: 19721004 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0904266106
- Primary Citation Related Structures:
3I1K, 3I1L, 3I26, 3I27 - PubMed Abstract:
Hemagglutinin esterases (HEs), closely related envelope glycoproteins in influenza C and corona- and toroviruses, mediate reversible attachment to O-acetylated sialic acids (Sias). They do so by acting both as lectins and as receptor-destroying enzymes, functions exerted by separate protein domains. HE divergence was accompanied by changes in quaternary structure and in receptor and substrate specificity. The selective forces underlying HE diversity and the molecular basis for Sia specificity are poorly understood. Here we present crystal structures of porcine and bovine torovirus HEs in complex with receptor analogs. Torovirus HEs form homodimers with sialate-O-acetylesterase domains almost identical to corresponding domains in orthomyxo- and coronavirus HEs, but with unique lectin sites. Structure-guided biochemical analysis of the esterase domains revealed that a functionally, but not structurally conserved arginine-Sia carboxylate interaction is critical for the binding and positioning of glycosidically bound Sias in the catalytic pocket. Although essential for efficient de-O-acetylation of Sias, this interaction is not required for catalysis nor does it affect substrate specificity. In fact, the distinct preference of the porcine torovirus enzyme for 9-mono- over 7,9-di-O-acetylated Sias can be explained from a single-residue difference with HEs of more promiscuous specificity. Apparently, esterase and lectin pockets coevolved; also the porcine torovirus HE receptor-binding site seems to have been designed to use 9-mono- and exclude di-O-acetylated Sias, possibly as an adaptation to replication in swine. Our findings shed light on HE evolution and provide fundamental insight into mechanisms of substrate binding, substrate recognition, and receptor selection in this important class of virion proteins.
- Virology Division, Department of Infectious Diseases & Immunology, Faculty of Veterinary Medicine, Utrecht University, 3584 CH Utrecht, The Netherlands.
Organizational Affiliation: