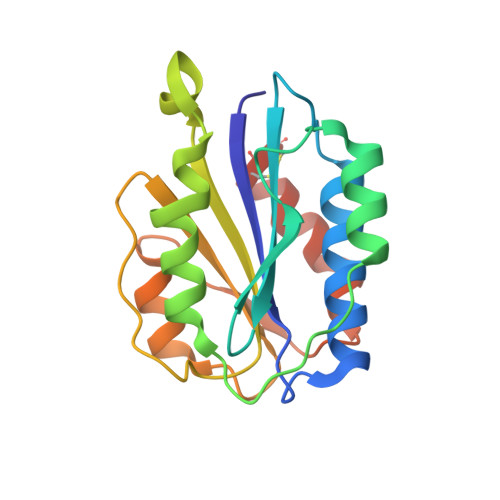
Structural specializations of A2, a force-sensing domain in the ultralarge vascular protein von Willebrand factor.
Zhang, Q., Zhou, Y.F., Zhang, C.Z., Zhang, X., Lu, C., Springer, T.A.(2009) Proc Natl Acad Sci U S A 106: 9226-9231
- PubMed: 19470641 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0903679106
- Primary Citation Related Structures:
3GXB - PubMed Abstract:
The lengths of von Willebrand factor (VWF) concatamers correlate with hemostatic potency. After secretion in plasma, length is regulated by hydrodynamic shear force-dependent unfolding of the A2 domain, which is then cleaved by a specific protease. The 1.9-A crystal structure of the A2 domain demonstrates evolutionary adaptations to this shear sensor function. Unique among VWF A (VWA) domains, A2 contains a loop in place of the alpha4 helix, and a cis-proline. The central beta4-strand is poorly packed, with multiple side-chain rotamers. The Tyr-Met cleavage site is buried in the beta4-strand in the central hydrophobic core, and the Tyr structurally links to the C-terminal alpha6-helix. The alpha6-helix ends in 2 Cys residues that are linked by an unusual vicinal disulfide bond that is buried in a hydrophobic pocket. These features may narrow the force range over which unfolding occurs and may also slow refolding. Von Willebrand disease mutations, which presumably lower the force at which A2 unfolds, are illuminated by the structure.
- Immune Disease Institute, Children's Hospital Boston and Department of Pathology, Harvard Medical School, Boston, MA 02115, USA.
Organizational Affiliation: