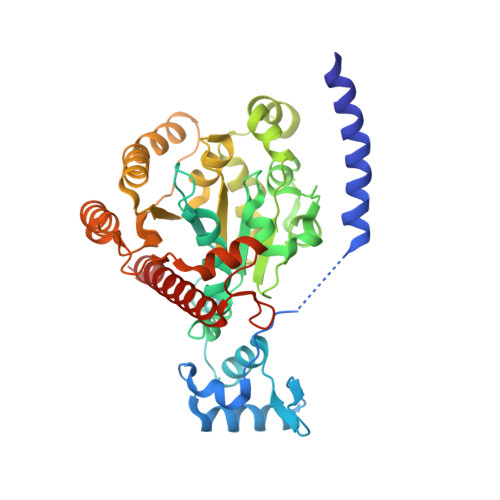
Structural basis for ADP-mediated transcriptional regulation by P1 and P7 ParA.
Dunham, T.D., Xu, W., Funnell, B.E., Schumacher, M.A.(2009) EMBO J 28: 1792-1802
- PubMed: 19461582 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/emboj.2009.120
- Primary Citation Related Structures:
3EZ2, 3EZ6, 3EZ7, 3EZ9, 3EZF - PubMed Abstract:
The accurate segregation of DNA is essential for the faithful inheritance of genetic information. Segregation of the prototypical P1 plasmid par system requires two proteins, ParA and ParB, and a centromere. When bound to ATP, ParA mediates segregation by interacting with centromere-bound ParB, but when bound to ADP, ParA fulfils a different function: DNA-binding transcription autoregulation. The structure of ParA is unknown as is how distinct nucleotides arbitrate its different functions. To address these questions, we carried out structural and biochemical studies. Crystal structures show that ParA consists of an elongated N-terminal alpha-helix, which unexpectedly mediates dimerization, a winged-HTH and a Walker-box containing C-domain. Biochemical data confirm that apoParA forms dimers at physiological concentrations. Comparisons of four apoParA structures reveal a strikingly flexible dimer interface that allows ParA to adopt multiple conformations. The ParA-ADP structure shows that ADP-binding activates DNA binding using a bipartite mechanism. First, it locks in one specific dimer conformation, and second, it induces the folding of two DNA-binding basic motifs that we show are critical for operator binding.
- Department of Biochemistry and Molecular Biology, MD Anderson Cancer Center, University of Texas, Houston, TX 77030, USA.
Organizational Affiliation: